University of Florida
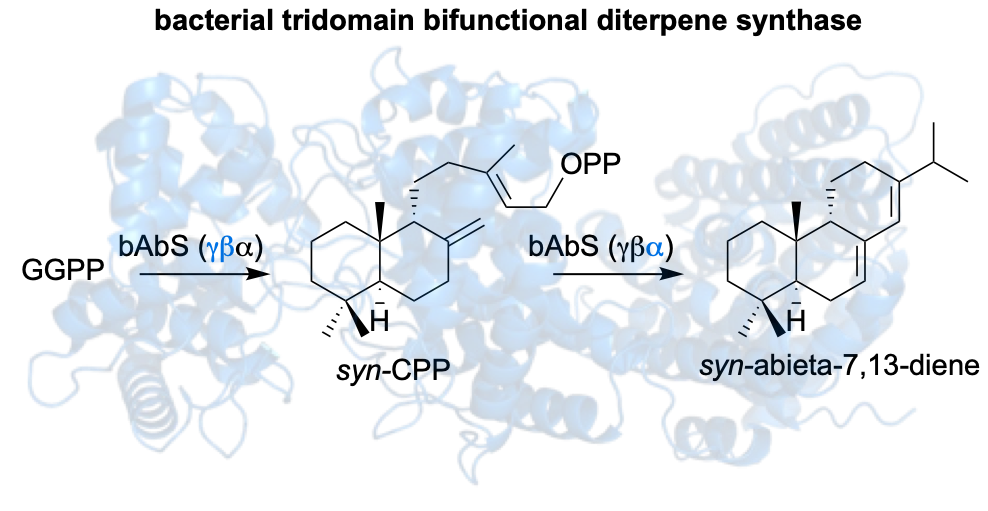
71. McCadden, C. A.; Łomowska-Keehner, D. P.; Qu, T.; Nafie, J.; Alsup, T. A.; Rudolf, J. D.* Discovery of a plant-like tridomain bifunctional syn-abieta-7,13-diene synthase in Streptomyces. Org. Biomol. Chem. 2025, doi: 10.1039/D5OB00724K.
Read the paper!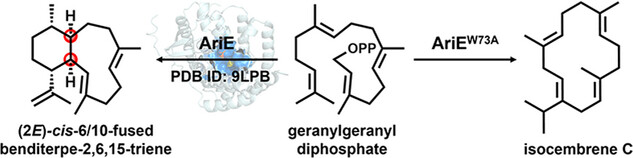
70. Li, F.-R.; Yang, Q.; He, J.; Sun, X.; Pan, X.; Xu, H.-M.; Rudolf, J. D.; Dong, L.-B.* Structural insights into the catalytic mechanism of cis-eunicellane cyclase AriE. Chem. Eur. J. 2025, 31, e202500012.
Read the paper!
69. Wei, X.; DeSnoo, W.; Li, Z.; Ning, W.; Kong, W.-Y.; Nafie, J.; Tantillo, D. J.;* Rudolf, J. D.* Avoidance of secondary carbocations, unusual deprotonation, and non-statistical dynamic effects in the cyclization mechanism of the 5/5/5/5-tetracyclic tetraisoquinane skeleton. J. Am. Chem. Soc. 2025, 147, 16293–16300.
Read the paper!
68. Wei, X.; Ning, W.; McCadden, C.; Alsup, T. A.; Li, Z.; Łomowska-Keehner, D. P.; Nafie, J.; Qu, T.; Osei Opoku, M.; Gillia, G. R.; Xu, B.; Icenhour, D. G.; Rudolf, J. D.* Exploring and expanding the natural chemical space of bacterial diterpenes. Nat. Commun. 2025, 16, 3721.
Read the paper!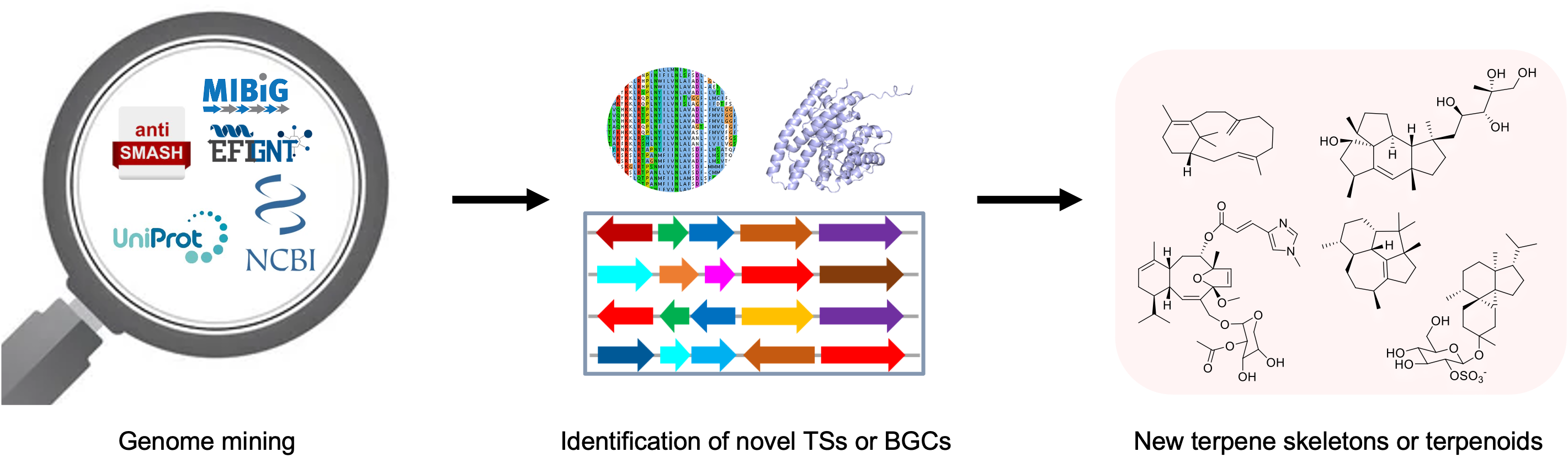
67. Ning, W.; Rudolf, J. D.* Discovery of bacterial terpenoids by genome mining. Meth. Enzymol. 2025, xxx, in press, doi: 10.1016/bs.mie.2025.01.078.
Read the paper!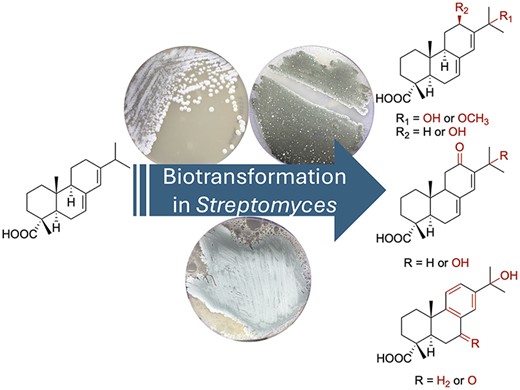
66. McCadden, C. A.; Alsup, T. A.; Ghiviriga, I.; Rudolf, J. D.* Biocatalytic diversification of abietic acid in Streptomyces. J. Ind. Microbiol. Biotechnol. 2025, 52, kuaf003.
Read the paper!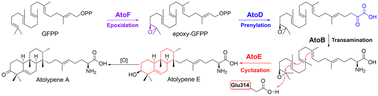
65. Wang, Z.; Alsup, T. A.; Pan, X.; Li, L.; Tian, J.; Yang, Z.; Lin, X.; Xu, H.-M.; Rudolf, J. D.; Dong, L.-B.* Biosynthesis of a bacterial meroterpenoid reveals a non-canonical class II meroterpenoid cyclase. Chem. Sci. 2025, 16, 310–317.
Read the paper!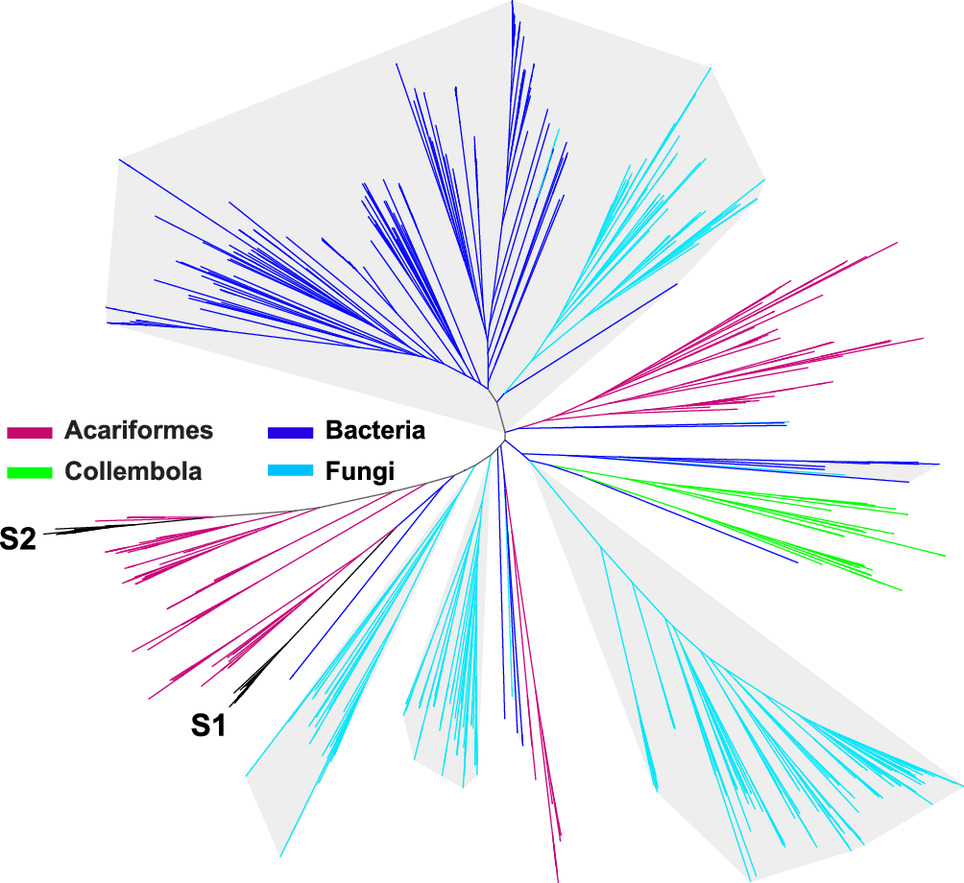
64. Chen, X.; Urban, J. M.; Wurlitzer, J.; Wei, X.; Han, J.; O’Connor, S. E.; Rudolf, J. D.; Köllner, T. G.; Chen, F. Canonical terpene synthases in arthropods: Intraphylum gene transfer. Proc. Natl. Acad. Sci. 2024, 121, e2413007121.
Read the paper!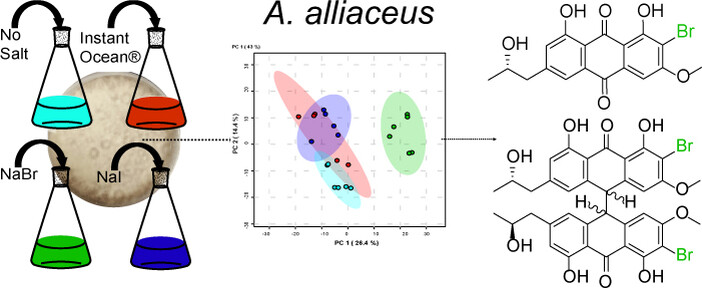
63. Mandelare, P. E.; Tee, S. S.; Icenhour, D. G.; Kaweesa, E. N.; McCauley, M.; Risinger, A. L.; Rudolf, J. D.; Loesgen, S.* Chemical diversity of Aspergillus alliaceus phenotypes: discovery of brominated bianthrones with activity against triple-negative breast cancer cell lines. ChemBioChem. 2024, 25, e202400398.
Read the paper!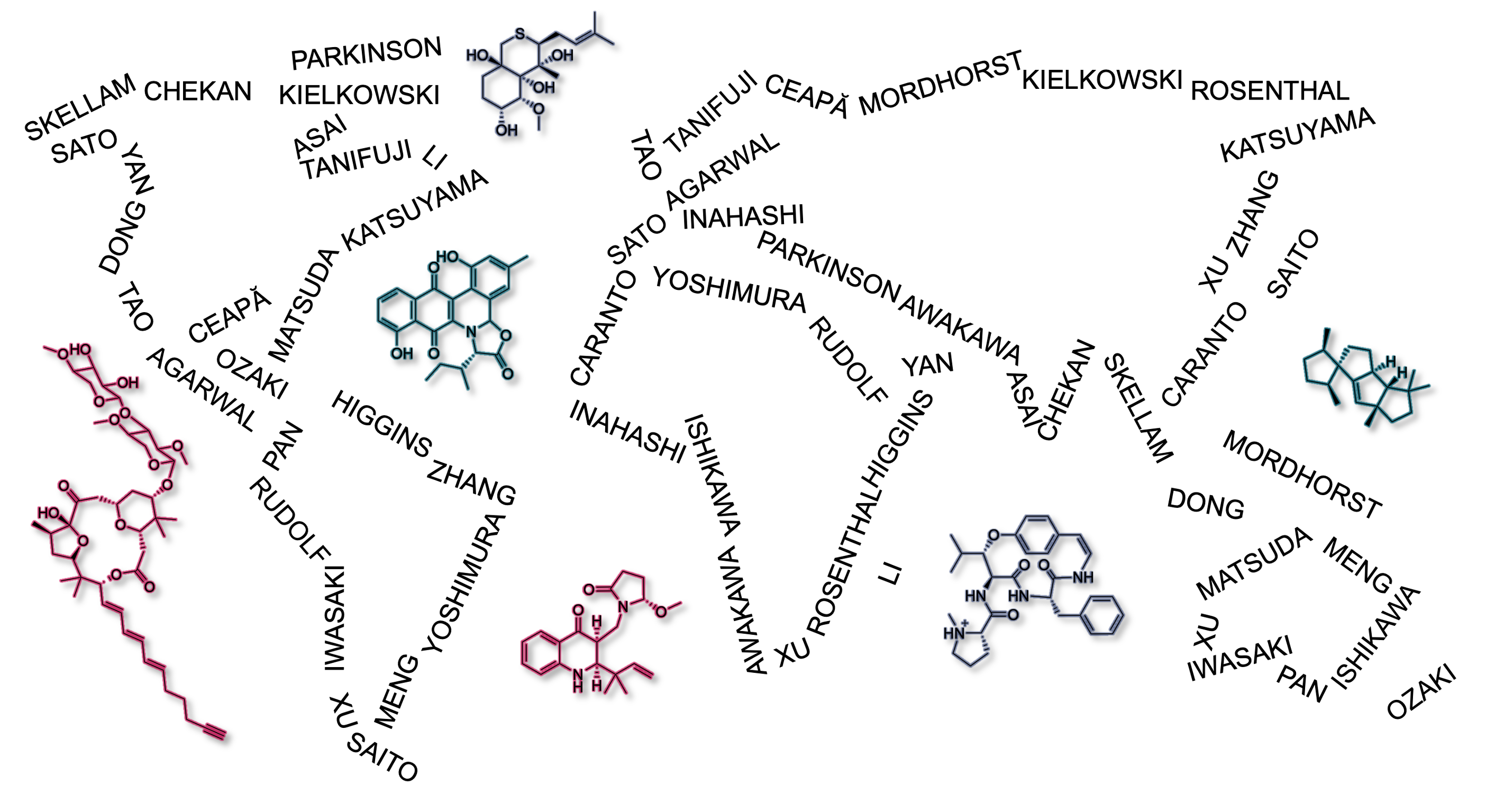
62. Rudolf, J. D.*; Barra, L.;* Awakawa, T.* Young investigators in natural products chemistry, biosynthesis, and enzymology. Beilstein J. Org. Chem. 2024, 20, 2720–2721. Special Issue Editorial.
Read the paper!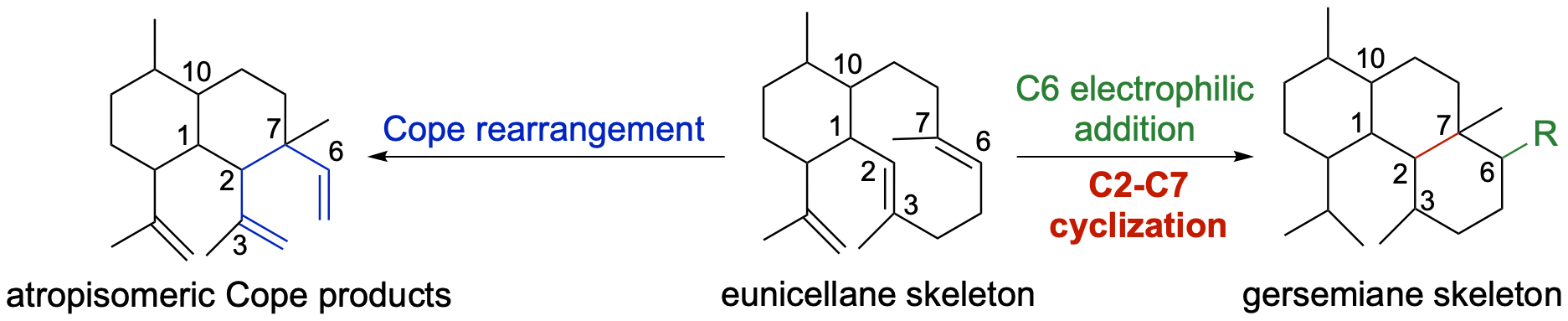
61. Li, Z.; Jindani, S.; Kojasoy, V.; Ortega, T.; Marshall, E.; Aboudd, K. A.; Loesgen, S.; Tantillo, D.;* Rudolf, J. D.* Computation-guided scaffold exploration of 2E,6E-1,10-trans/cis eunicellanes. Beilstein J. Org. Chem. 2024, 20, 1320–1326.
Read the paper!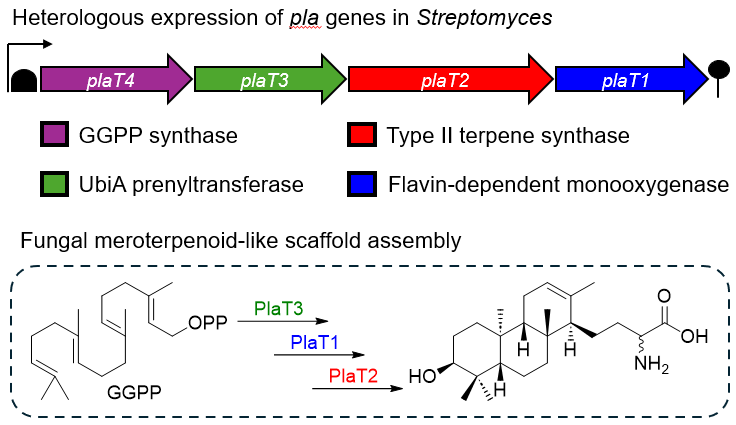
60. Alsup, T. A.; Li, Z.; McCadden, C.; Jagels, A.; Łomowska-Keehner, D. P.; Marshall, E. M.; Dong, L.-B.; Loesgen, S.; Rudolf, J. D.* Early-stage biosynthesis of phenalinolactone diterpenoids involves sequential prenylation, epoxidation, and cyclization. RSC Chem. Biol. 2024, 5, 1010–1016.
Read the paper!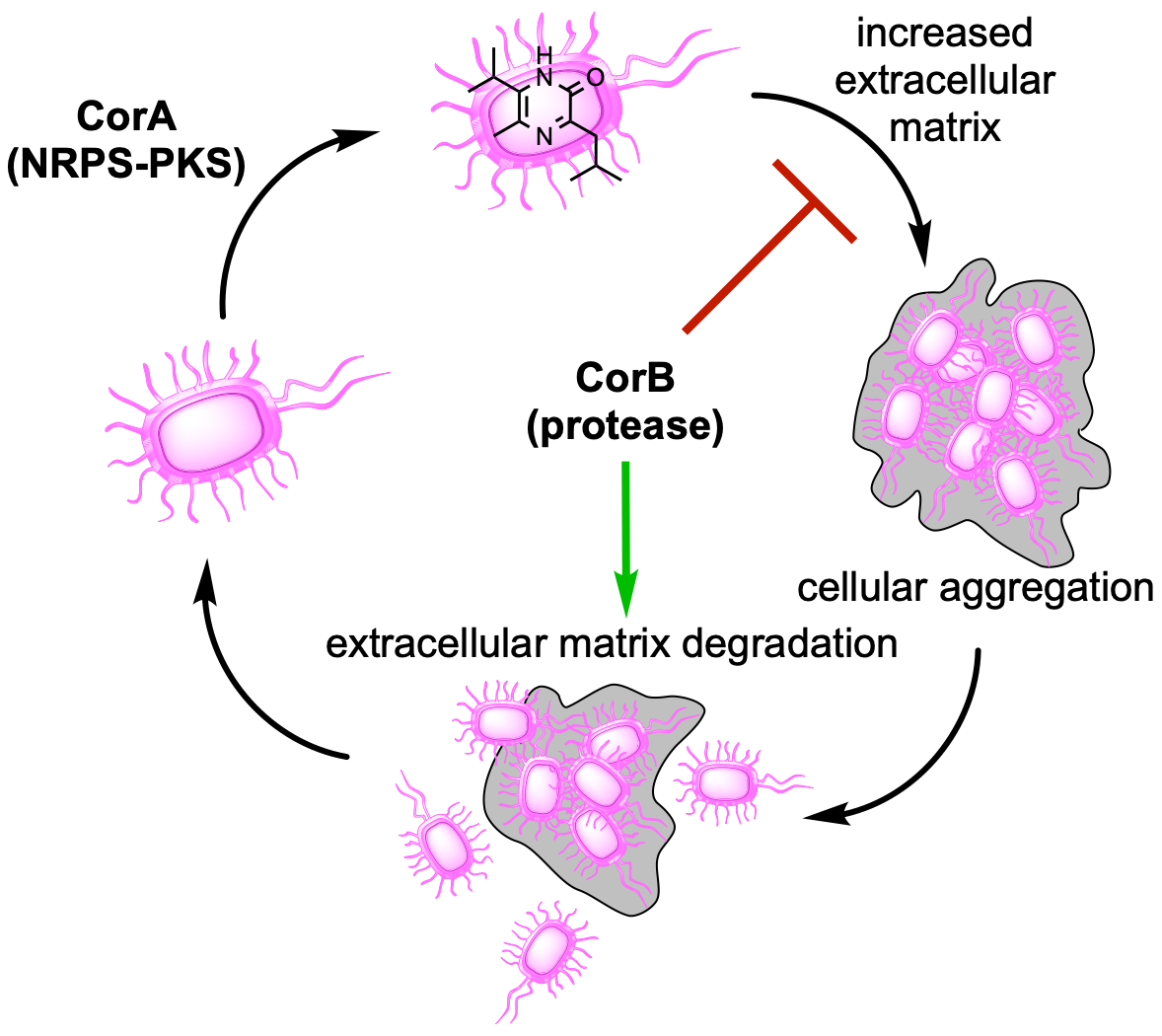
59. Rudolf, J. D.*; Loesgen, S.* Pyrazinone biosynthesis and signaling – myxostyle. ACS Cent. Sci. 2024, 10, 511–513. Invited First Reaction.
Read the paper!
58. Rudolf, J. D.* Preface to Terpene Synthases. Meth. Enzymol. 2024, 699, xxi–xxiii.
Read the paper!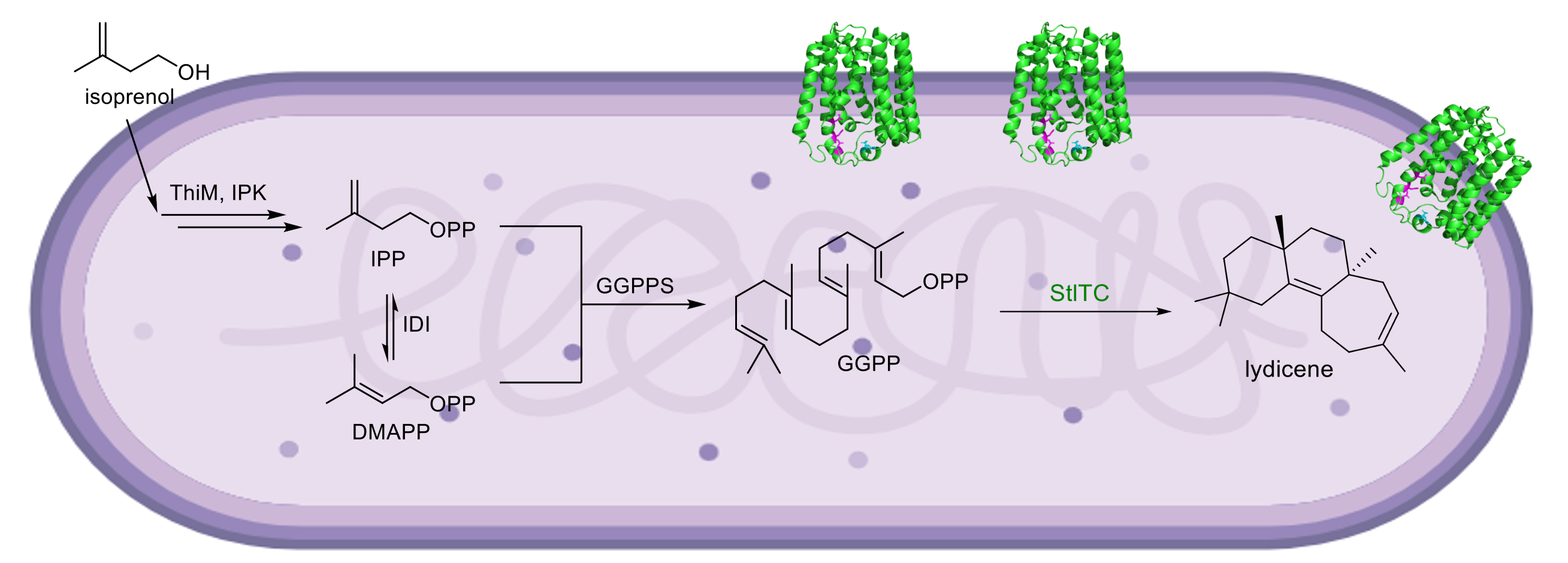
57. Alsup, T. A.; Osei Opoku, M.; Rudolf, J. D.* Characterization of UbiA terpene synthases with a precursor overproduction system in Escherichia coli. Meth. Enzymol. 2024, 699, 395–417.
Read the paper!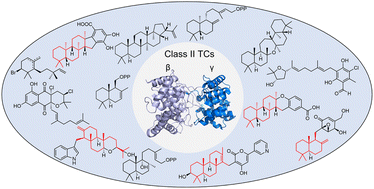
55. Pan, X.; Rudolf, J. D.;* Dong, L.-B.* Class II terpene cyclases: structures, mechanisms, and engineering. Nat. Prod. Rep. 2024, 41, 402–433.
Read the paper!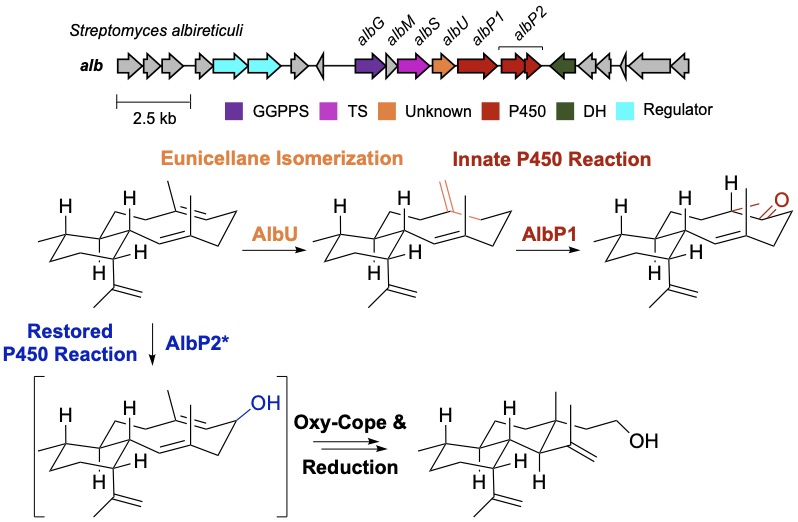
54. Li, Z.; Xu, B.; Alsup, T. A.; Wei, X.; Ning, W.; Icenhour, D. G.; Ehrenberger, M. A.; Ghiviriga, I.; Giang, B.-D.; Rudolf, J. D.* Cryptic isomerization in diterpene biosynthesis and the restoration of an evolutionarily defunct P450. J. Am. Chem. Soc. 2023, 145, 22361–22365.
Read the paper!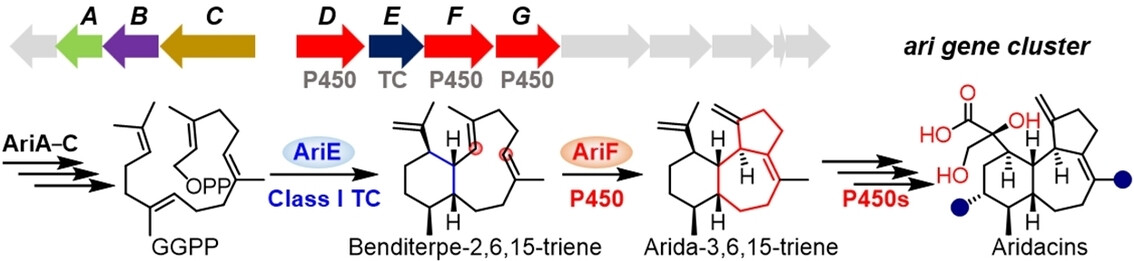
53. Wang, Z.; Yang, Q.; He, J.; Li, H.; Pan, X.; Li, Z.; Xu, H.-M.; Rudolf, J. D.; Tantillo, D. J.*; Dong, L.-B.* Cytochrome P450-mediated cyclization in eunicellane-derived diterpenoid biosynthesis. Angew. Chem. Int. Ed. 2023, 62, e202312490.
Read the paper!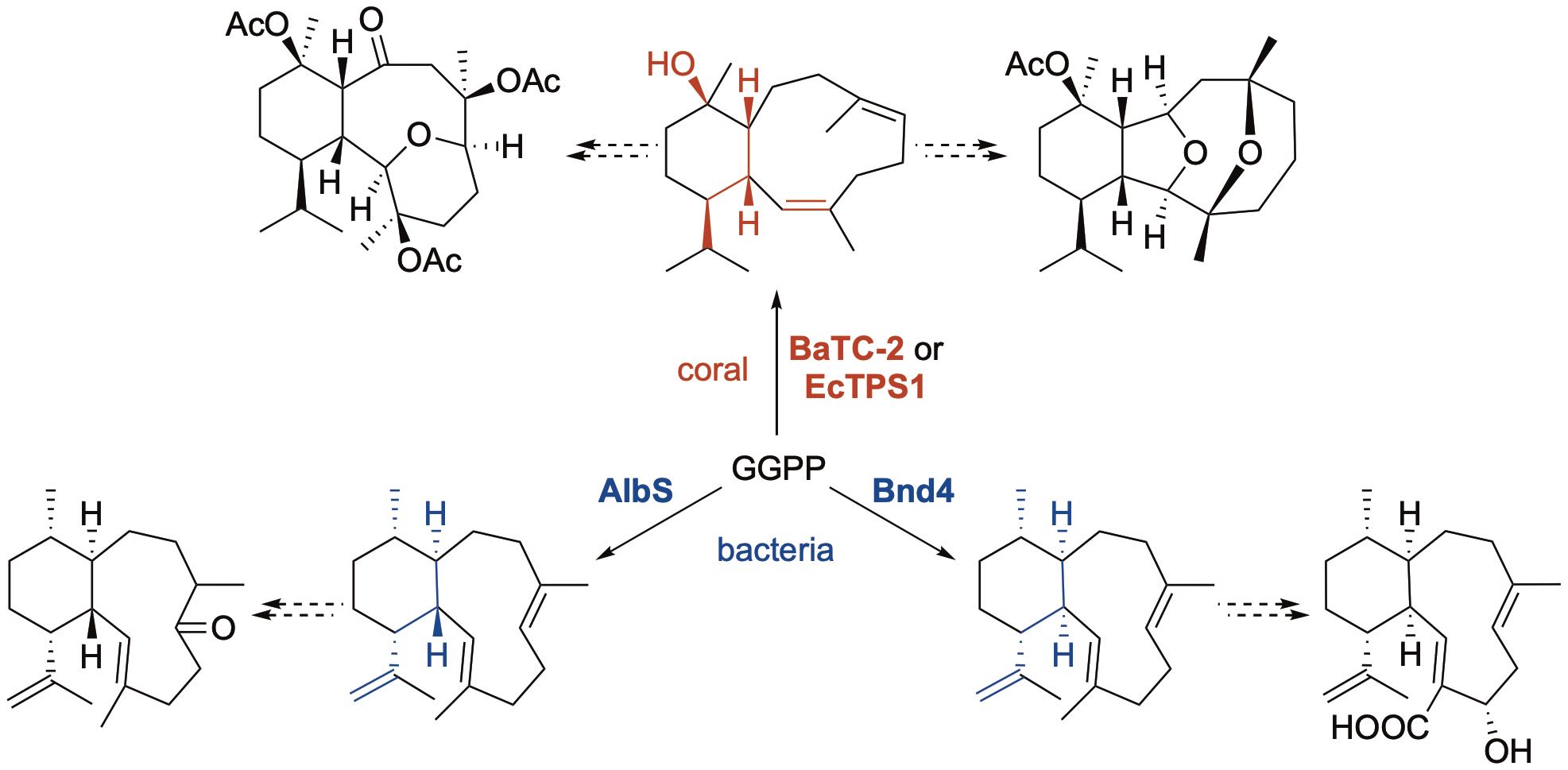
52. Li, Z.; Rudolf, J. D.* Biosynthesis, enzymology, and future of eunicellane diterpenoids. J. Ind. Microbiol. Biotechnol. 2023, 50, kuad027.
Read the paper!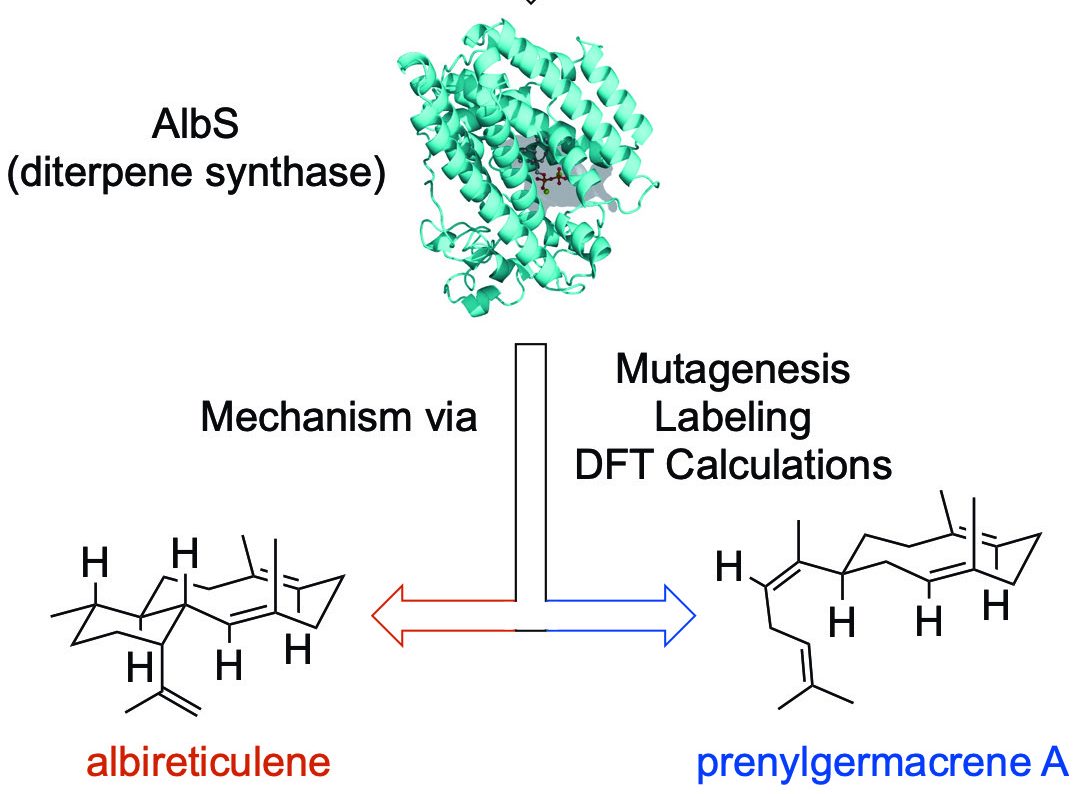
51. Li, Z.; Xu, B.; Kojasoy, V.; Ortega, T.; Adpressa, D.; Ning, W.; Wei, X.; Liu, J.; Tantillo, D. J.; Loesgen, S.; Rudolf, J. D.* First trans-eunicellane terpene synthase in bacteria. Chem 2023, 9, 698–708.
Read the paper!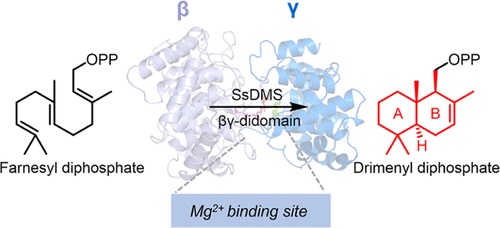
50. Pan, X.; Du, W.; Zhang, X.; Lin, X.; Li, F.-R.; Yang, Q.; Wang, H.; Rudolf, J. D.;* Zhang, B.;* Dong, L.-B.* Discovery, structure, and mechanism of a class II sesquiterpene cyclase. J. Am. Chem. Soc. 2022, 144, 22067–22074.
Read the paper!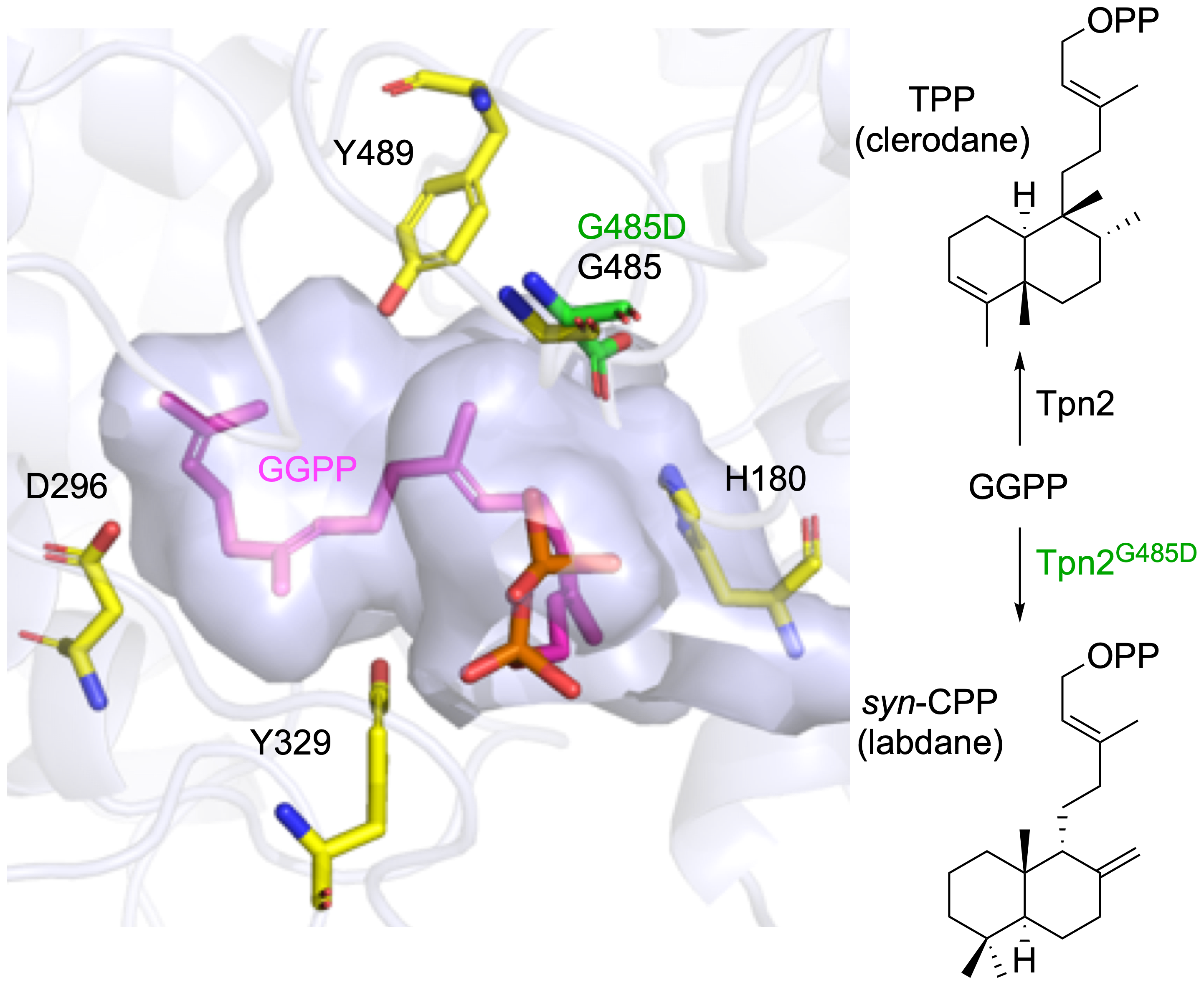
49. Stowell, E. A.; Ehrenberger, M. E.; Lin, Y.-L.; Chang, C.-Y.;* Rudolf, J. D.* Structure-guided product determination of the bacterial type II diterpene synthase Tpn2. Commun. Chem. 2022, 5, 146.
Read the paper!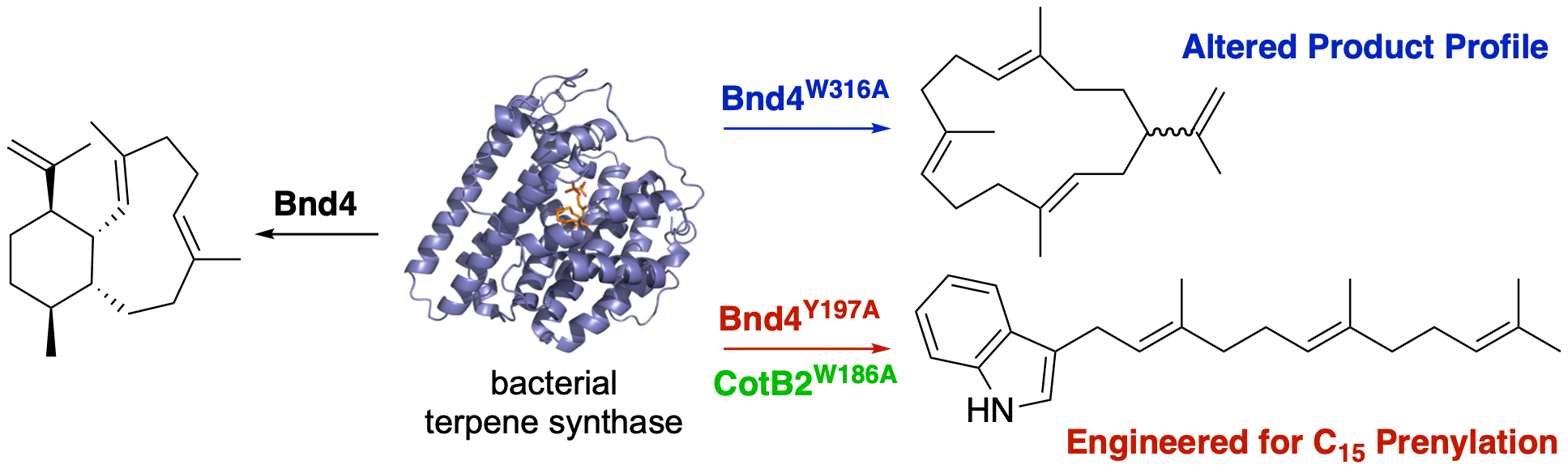
48. Xu, B.; Ning, W.; Wei, X.; Rudolf, J. D.* Mutation of the eunicellane synthase Bnd4 alters its product profile and expands its prenylation ability. Org. Biomol. Chem. 2022, 20, 8833–8837.
Read the paper!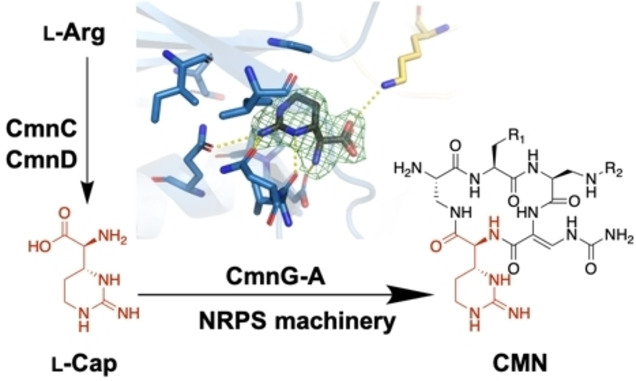
47. Chen, I.-H.; Cheng, T.; Wang, Y.-L.; Huang, S.-J.; Hsiao, Y.-H.; Lai, Y.-T.; Rudolf, J. D.; Chang, C.-Y.* Characterization and structural determination of CmnG-A, the adenylation domain that activates the nonproteinogenic amino acid capreomycidine in capreomycin biosynthesis. ChemBioChem 2022, 23, e202200563.
Read the paper!
46. de Crecy-Lagard, V.;* […] Rudolf, J. D.; […] (58 total authors). A roadmap for the functional annotation of protein families: a community perspective. Database 2022, baac062.
Read the paper!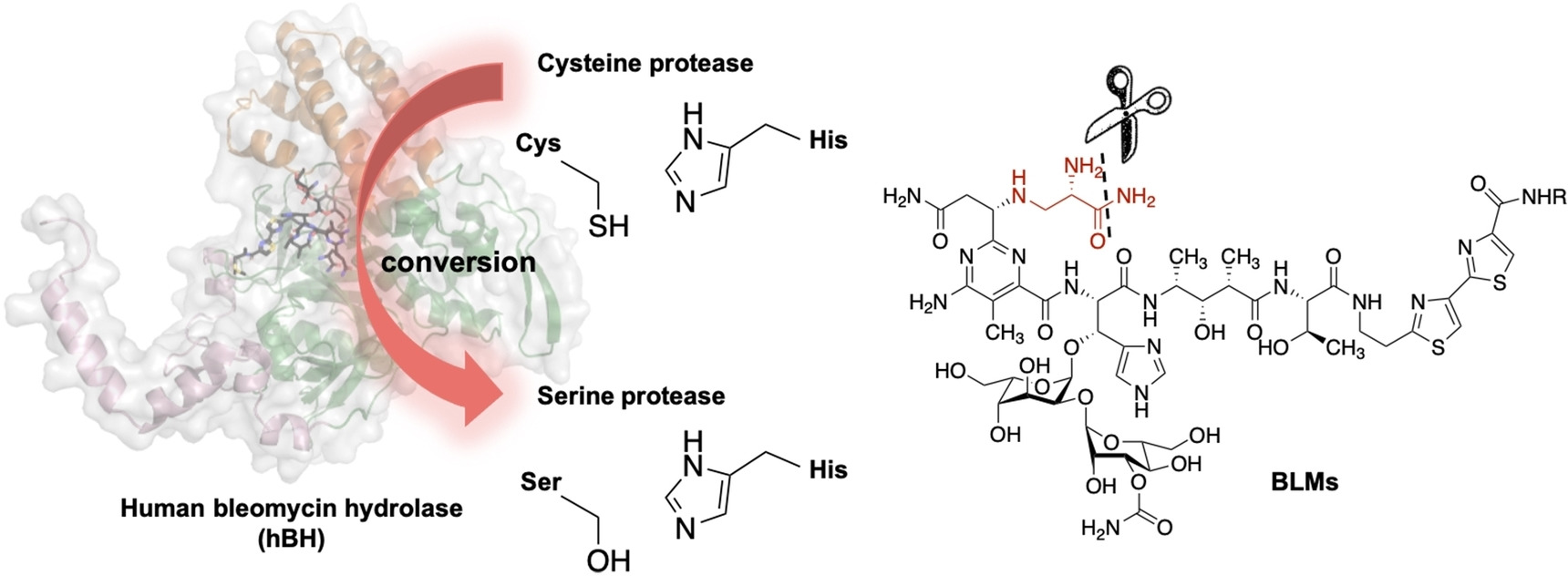
45. Zheng, Y.-Z.; Cui, J.; Wang, Y.-L.; Huang, S.-J.; Lin, E.-C.; Huang, S.-C.; Rudolf, J. D.; Yan, X.; Chang, C.-Y.* The structure-function relationship of human bleomycin hydrolase: mutation of a cysteine protease into a serine protease. ChemBioChem 2022, 23, e202200186.
Read the paper!
44. van Santen, J. A.; Poynton, E. F.; Iskakova, D.; McMann, E.; Alsup, T. A.; Clark, T. N.; Fergusson, C. H.; Fewer, D. P.; Hughes, A. H.; McCadden, C. A.; Parra, J.; Soldatou, S.; Rudolf, J. D.; Janssen, E. M-L.; Duncan, K. R.; Linington, R. G.* The Natural Products Atlas 2.0: a database of microbially-derived natural products. Nucl. Acids Res. 2022, 50, D1317–D1323.
Read the paper!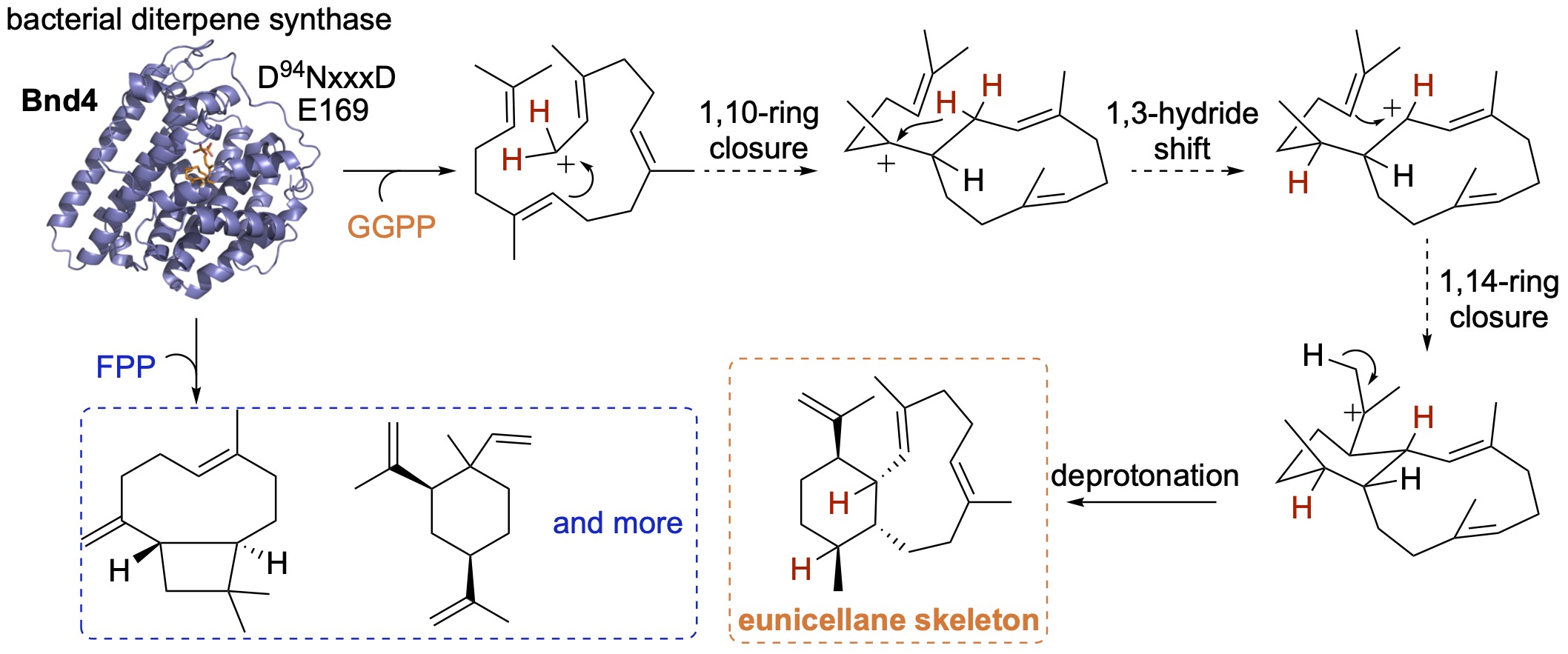
43. Xu, B.; Tantillo, D. J.; Rudolf, J. D.* Mechanistic insights into the formation of the 6,10-bicyclic eunicellane skeleton by the bacterial diterpene synthase Bnd4. Angew. Chem. Int. Ed. 2021, 60, 23159–23163.
Read the paper!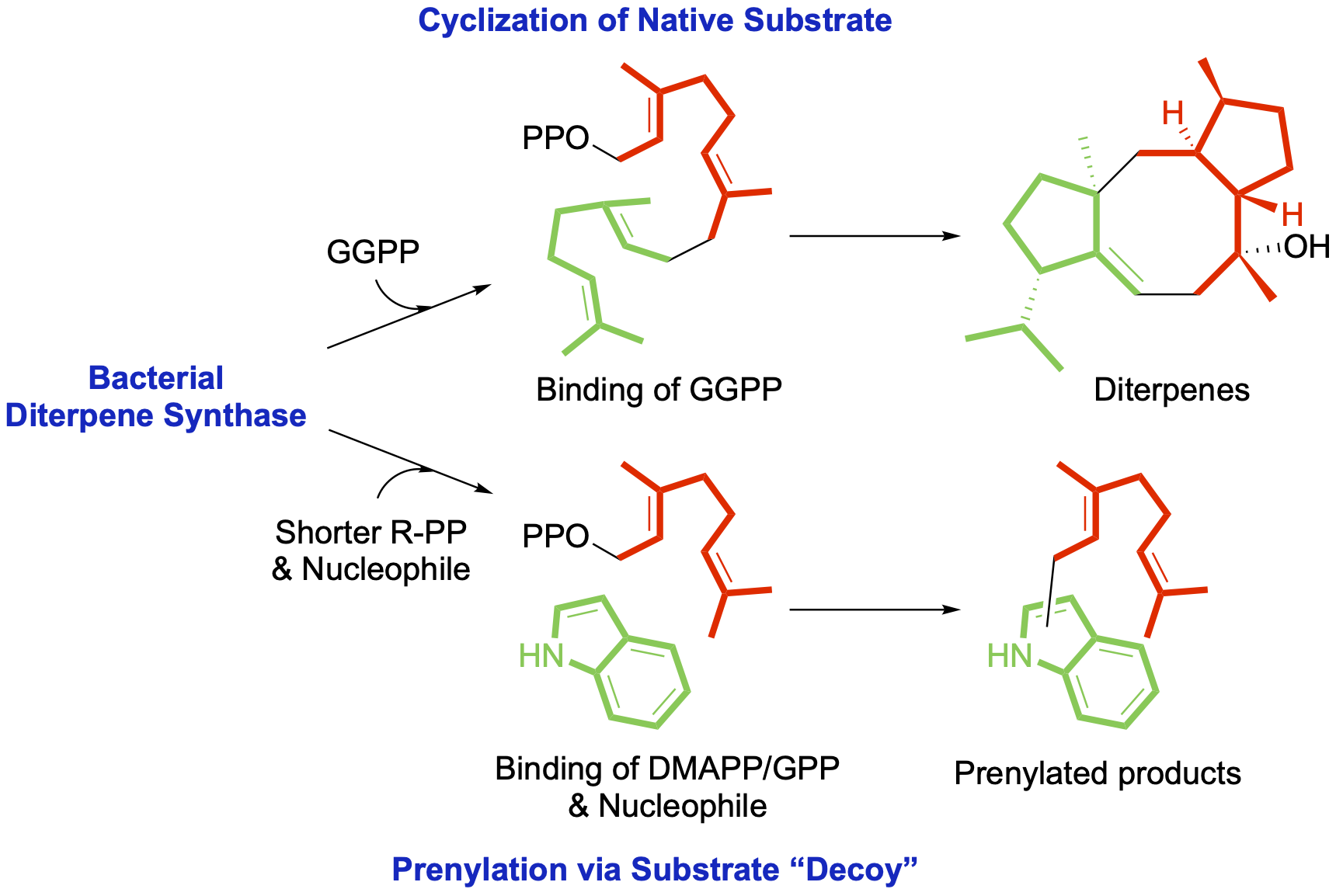
42. Xu, B.; Li, Z.; Alsup, T. A.; Ehrenberger, M. A.; Rudolf, J. D.* Bacterial diterpene synthases prenylate small molecules. ACS Catal. 2021, 11, 5906–5915.
Read the paper!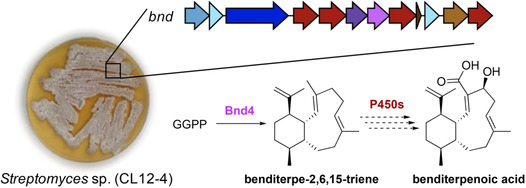
41. Zhu, C.; Xu, B.; Address, D. A.; Rudolf, J. D.;* Loesgen, S.* Discovery and biosynthesis of a structurally dynamic antibacterial diterpenoid. Angew. Chem. Int. Ed. 2021, 60, 14163–14170.
Read the paper!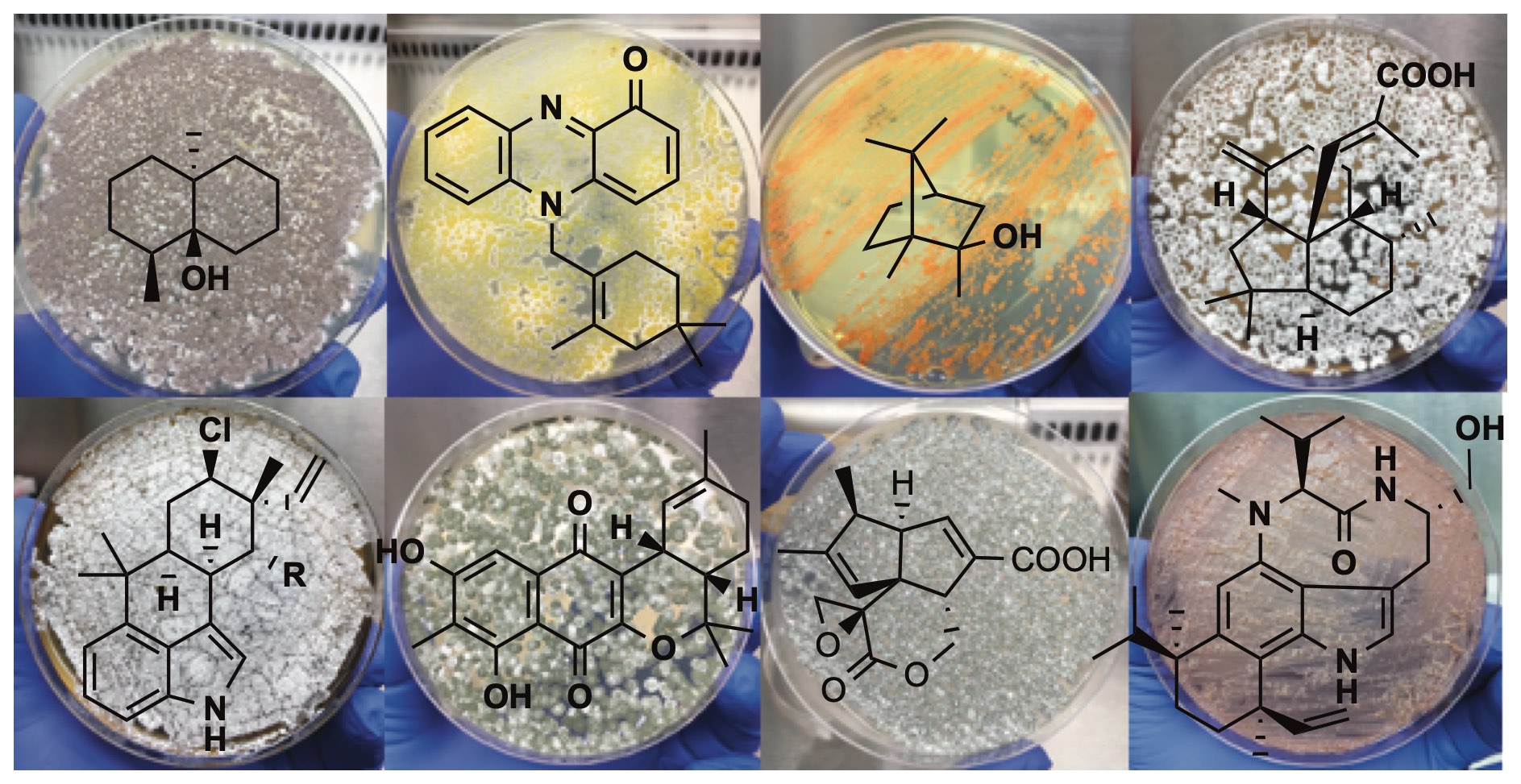
40. Rudolf, J. D.;* Alsup, T. A.; Xu, B.; Li, Z. Bacterial terpenome. Nat. Prod. Rep. 2021, 38, 905–980.
Read the paper!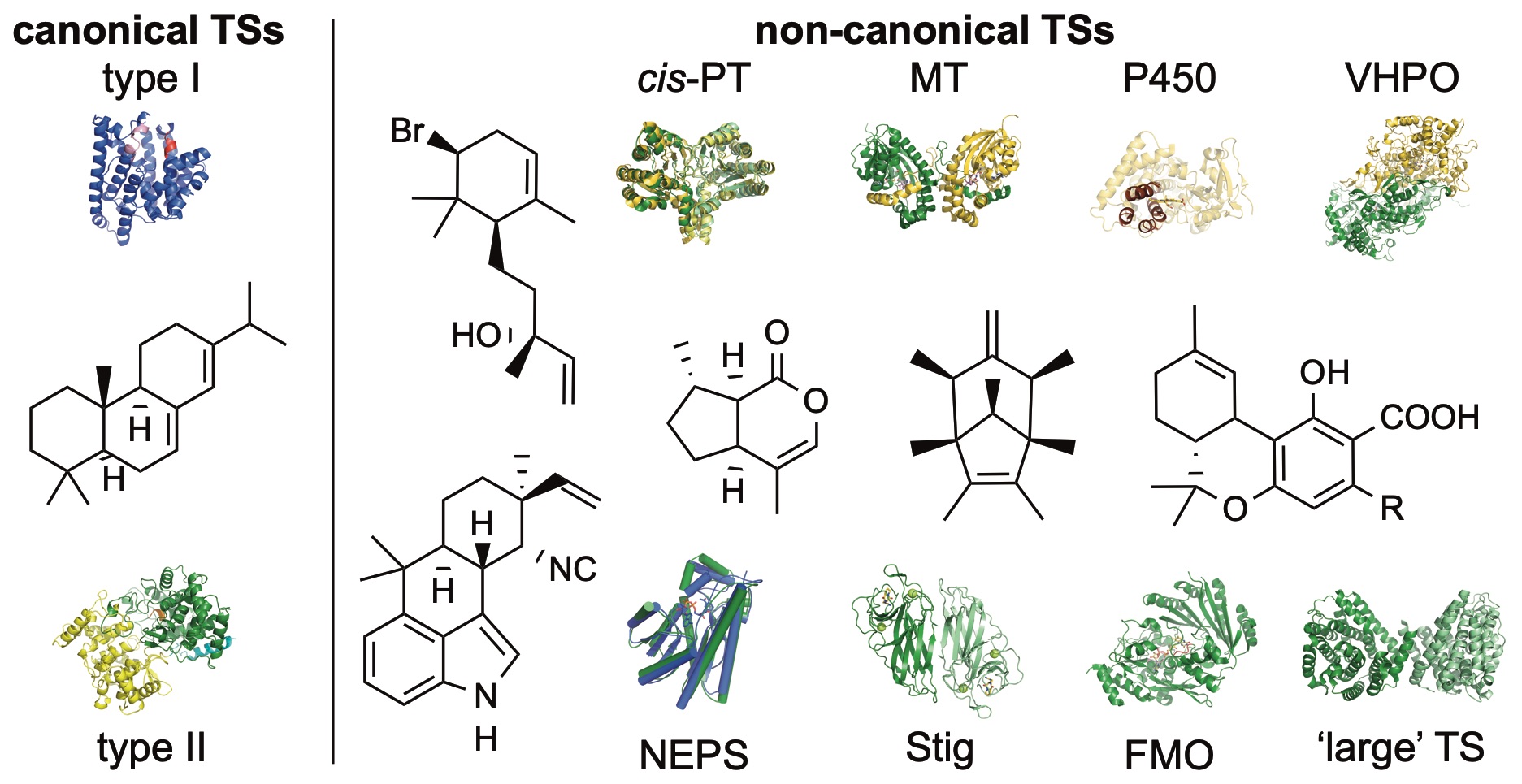
38. Rudolf, J. D.;* Chang, C.-Y. Terpene synthases in disguise: enzymology, structure, and opportunities of non-canonical terpene synthases. Nat. Prod. Rep. 2020, 37, 425–463.
Read the paper!Scripps Florida – Postdoc
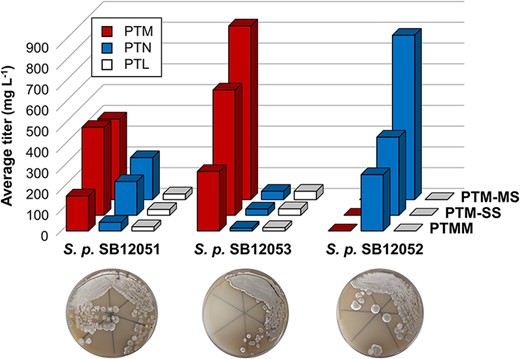
56. Fluegel, L. L.; Deng, M.-R.; Su, P.; Kalkreuter, E.; Yang, D.; Rudolf, J. D.; Dong, L.-B.; Shen, B.* Development of platensimycin, platencin, and platensilin overproducers by biosynthetic pathway engineering and fermentation medium optimization. J. Ind. Microbiol. Biotechnology. 2024, 51, kuae003.
Read the paper!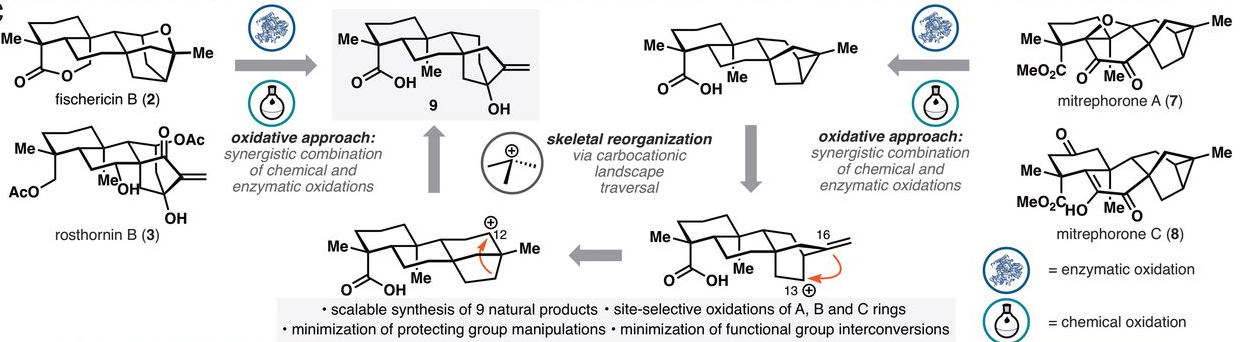
39. Zhang, X.; King-Smith, E.; Dong, L.-B.; Yang, L.-C.; Rudolf, J. D.; Shen, B.; Renata, H.* Divergent synthesis of complex diterpenes through a hybrid oxidative approach. Science 2020, 369, 799–806.
Read the paper!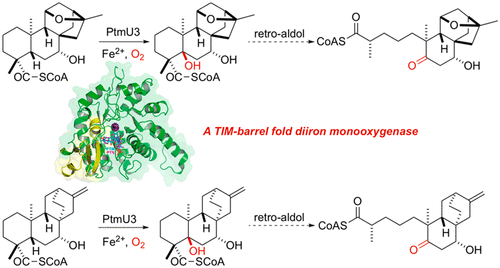
37. Dong, L.-B.; Liu, Y.-C.; Cepeda, A.; Kalkreuter, E.; Deng, M.-R.; Rudolf, J. D.; Chang, C.; Joachimiak, A.; Phillips Jr., G. N.; Shen, B.* Characterization and crystal structure of a nonheme diiron monooxygenase involved in platensimycin and platencin biosynthesis. J. Am. Chem. Soc. 2019, 141, 12406–12412.
Read the paper!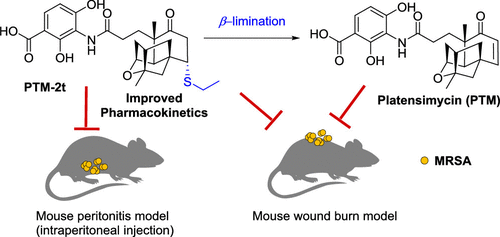
36. Su, M.; Qiu, L.; Deng, Y.; Ruiz, C. H.; Rudolf, J. D.; Dong, L.-B.; Feng, X.; Cameron, M. D.; Shen, B.; Duan, Y.; Huang, Y.* Evaluation of platensimycin and platensimycin-inspired thioether analogues against methicillin-resistant Staphylococcus aureus in topical and systemic infection mouse models. Mol. Pharm. 2019, 16, 3065–3071.
Read the paper!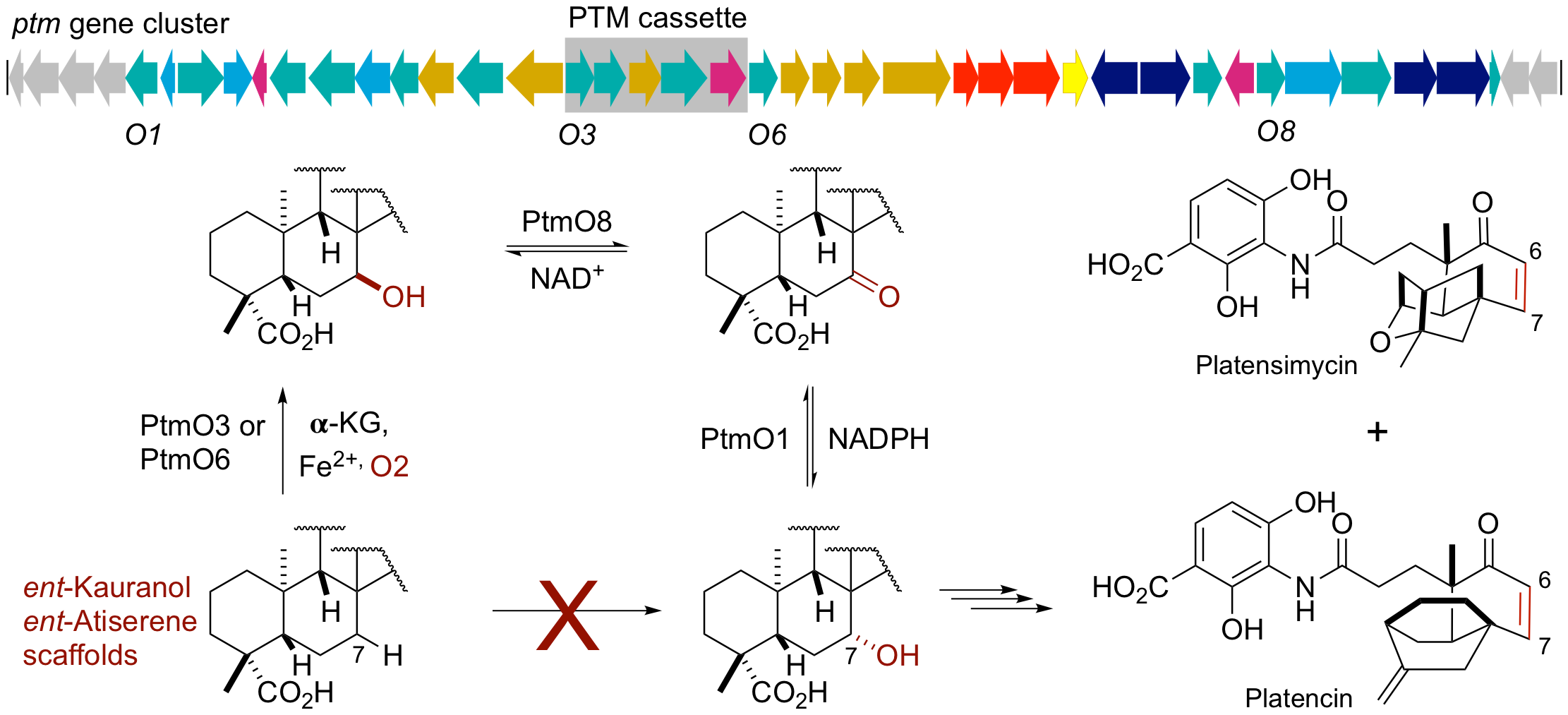
35. Dong, L.-B.; Zhang, X.; Rudolf, J. D.; Deng, M.-R.; Kalkreuter, E.; Cepeda, A. J.; Renata, H.; Shen, B.* Cryptic and stereospecific hydroxylation, oxidation, and reduction in platensimycin and platencin biosynthesis. J. Am. Chem. Soc. 2019, 141, 4043–4050.
Read the paper!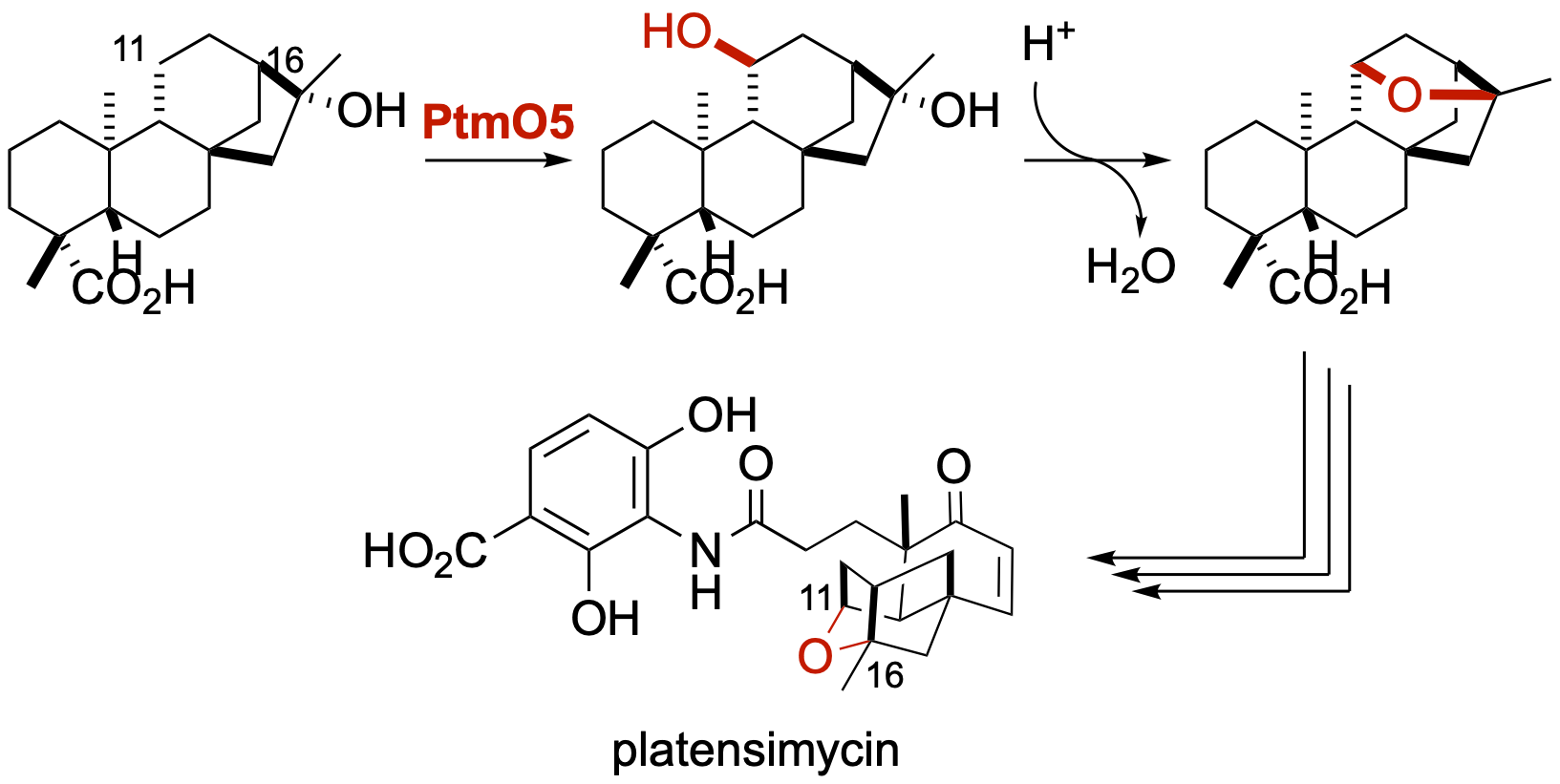
34. Rudolf, J. D.; Dong, L.-B.; Zhang, X.; Renata, H.; Shen, B.* Cytochrome P450-catalyzed hydroxylation initiating ether formation in platensimycin biosynthesis. J. Am. Chem. Soc. 2018, 140, 12349–12353.
Read the paper!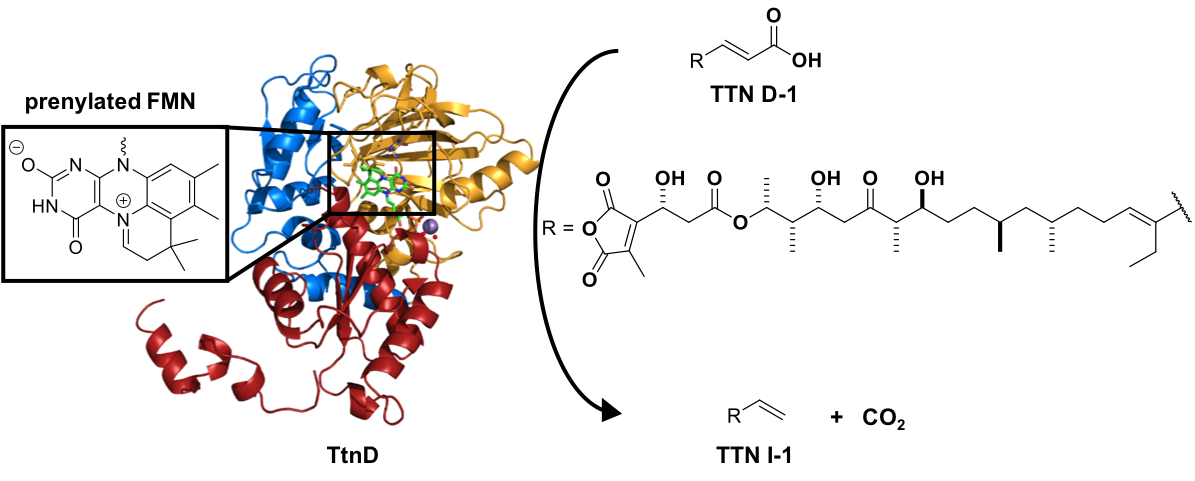
33. Annaval, T.; Han, L.; Rudolf, J. D.; Xie G.; Yang, D.; Chang, C.-Y.; Ma, M.; Crnovcic, I.; Miller, M. D.; Soman, J.; Xu, W.; Phillips Jr., G. N.; Shen, B.* Biochemical and structural characterization of TtnD, a prenylated FMN-dependent decarboxylase from the tautomycetin biosynthetic pathway. ACS Chem. Biol. 2018, 13, 2728–2738.
Read the paper!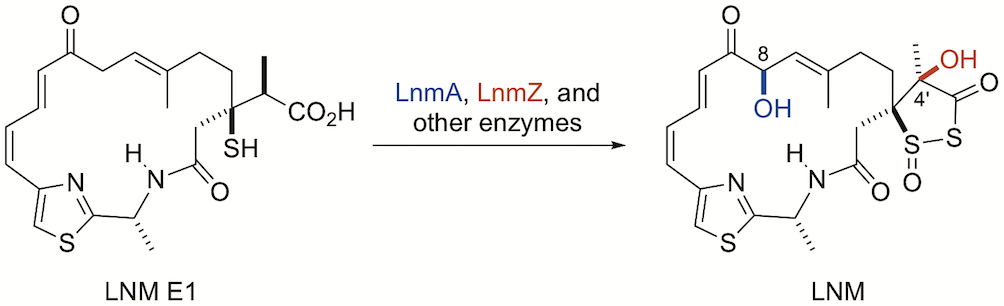
32. Kwong, T.; Ma, M.; Pan, G.; Hindra; Yang, D.; Lohman, J. R..; Rudolf, J. D.; Cleveland, J. L.; Shen, B.* P450-catalyzed tailoring steps in leinamycin biosynthesis featuring regio- and stereoselectivity hydroxylations and substrate promiscuities. Biochemistry 2018, 57, 5005–5013.
Read the paper!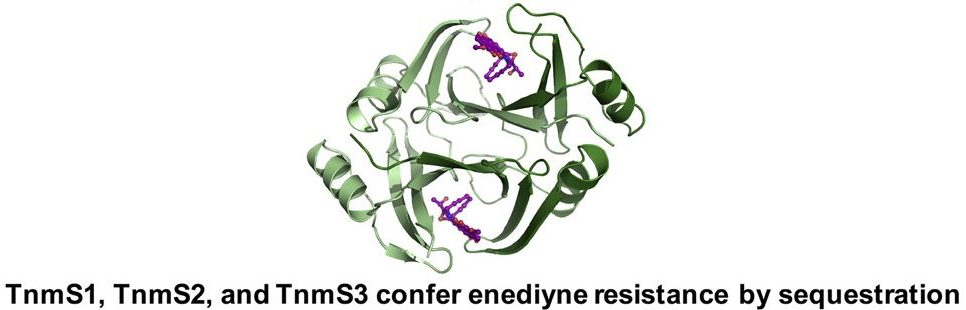
31. Chang, C.-Y.; Yan, X.; Crnovcic, I.; Annaval, T.; Chang, C.; Nocek, B.; Rudolf, J. D.; Yang, D.; Hindra; Babnigg, G.; Joachimiak, A.; Phillips Jr., G. N.; Shen, B.* Resistance to enediyne antitumor antibiotics by sequestration. Cell Chem. Biol. 2018, 25, 1075–1085.
Read the paper!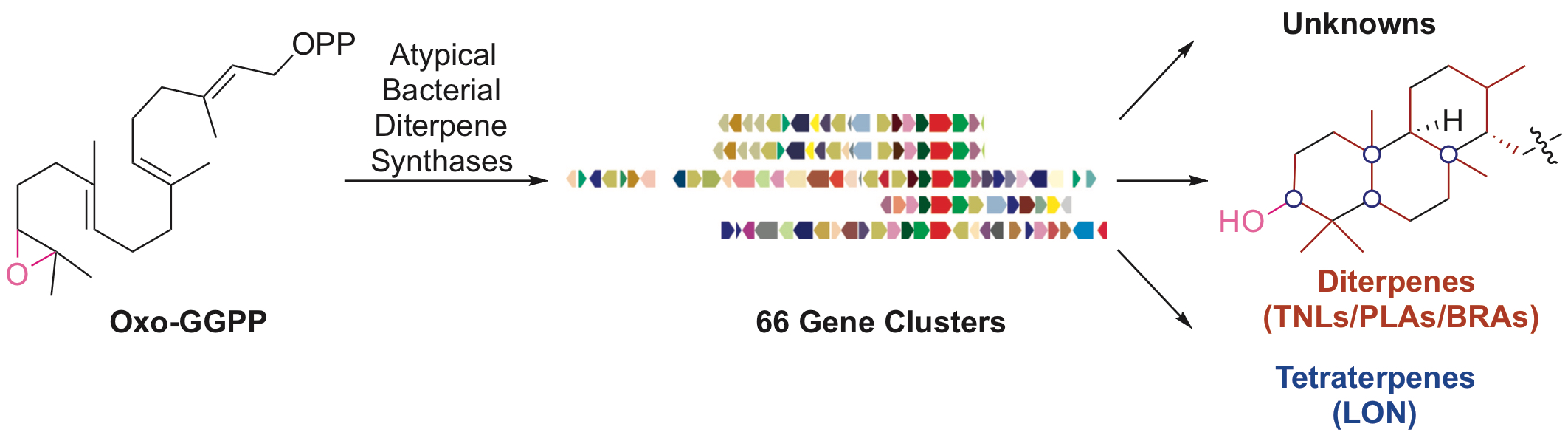
30. Dong, L.-B.; Rudolf, J. D.; Deng, M.-R.; Yan, Y.; Shen, B.* Discovery of the tiancilactone antibiotics by genome mining of atypical bacterial type II diterpene synthases. ChemBioChem 2018, 19, 1727–1733.
Read the paper!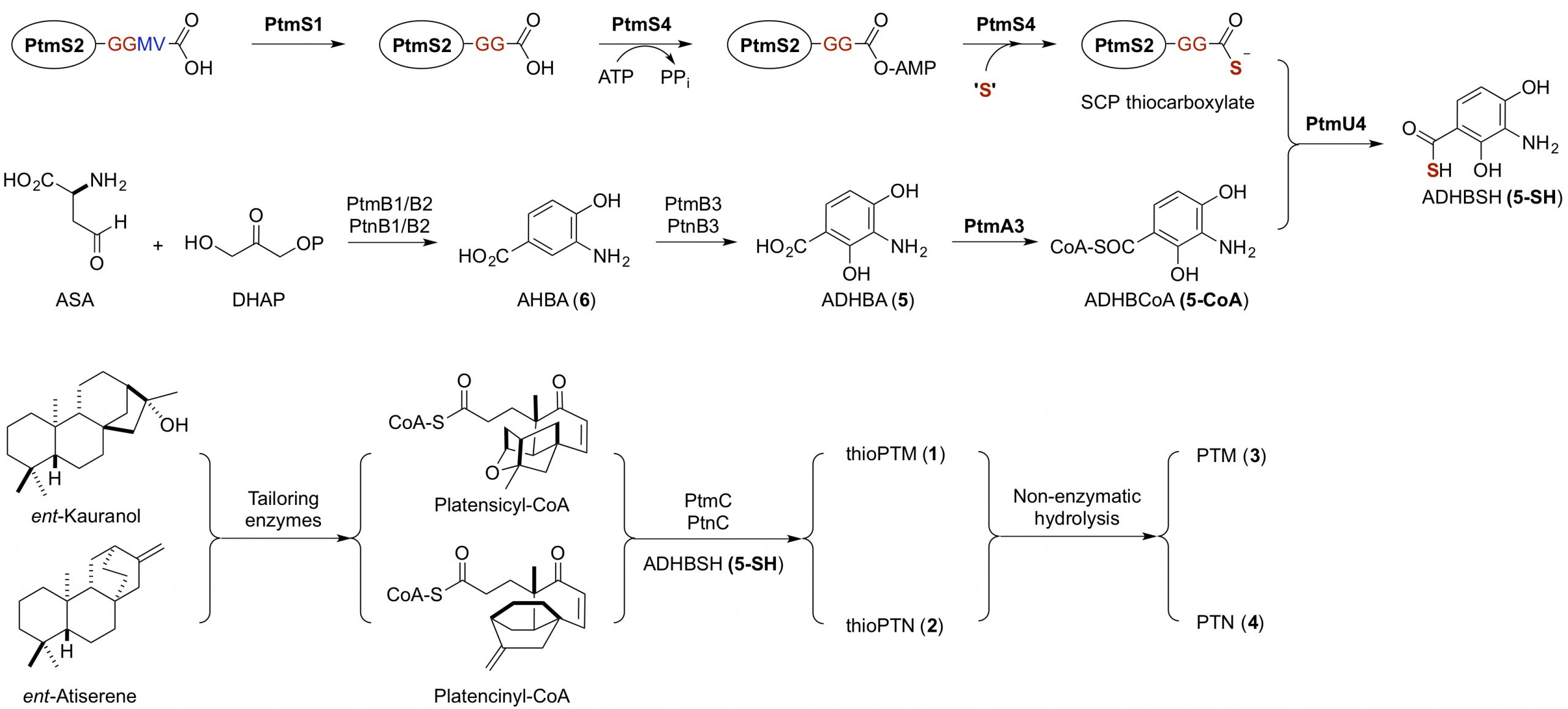
29. Dong, L.-B.; Rudolf, J. D.; Kang, D.; Wang, N.; He, C. Q.; Deng, Y.; Huang, Y.; Houk, K. N.; Duan, Y.; Shen, B.* Biosynthesis of thiocarboxylic acid-containing natural products. Nat. Commun. 2018, 9, 2362.
Read the paper!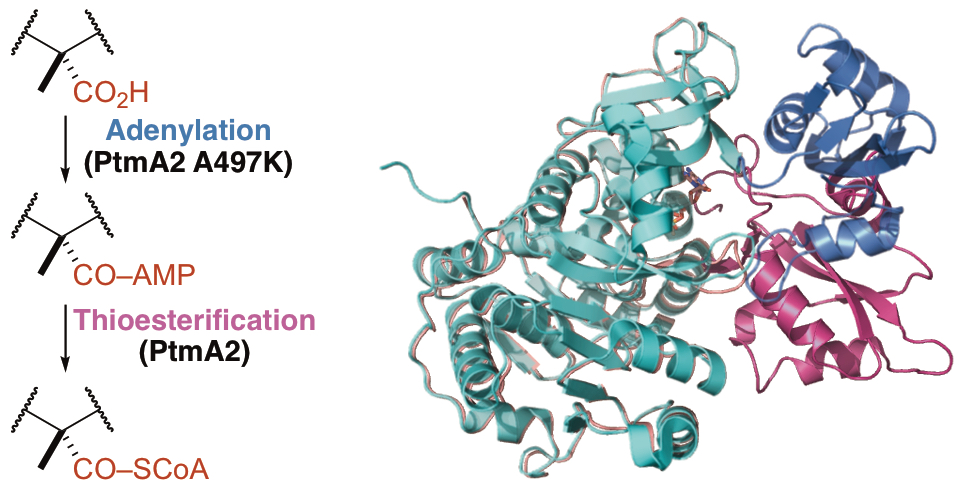
28. Wang, N.; Rudolf, J. D.; Dong, L.-B.; Osipiuk, J.; Hatzos-Skintges, C.; Endres, M.; Chang, C.-Y.; Babnigg, G.; Joachimiak, A.; Phillips Jr., G. N.; Shen, B.* Natural separation of the acyl-CoA ligase reaction results in a non-adenylating enzyme. Nat. Chem. Biol. 2018, 14, 730–737.
Read the paper!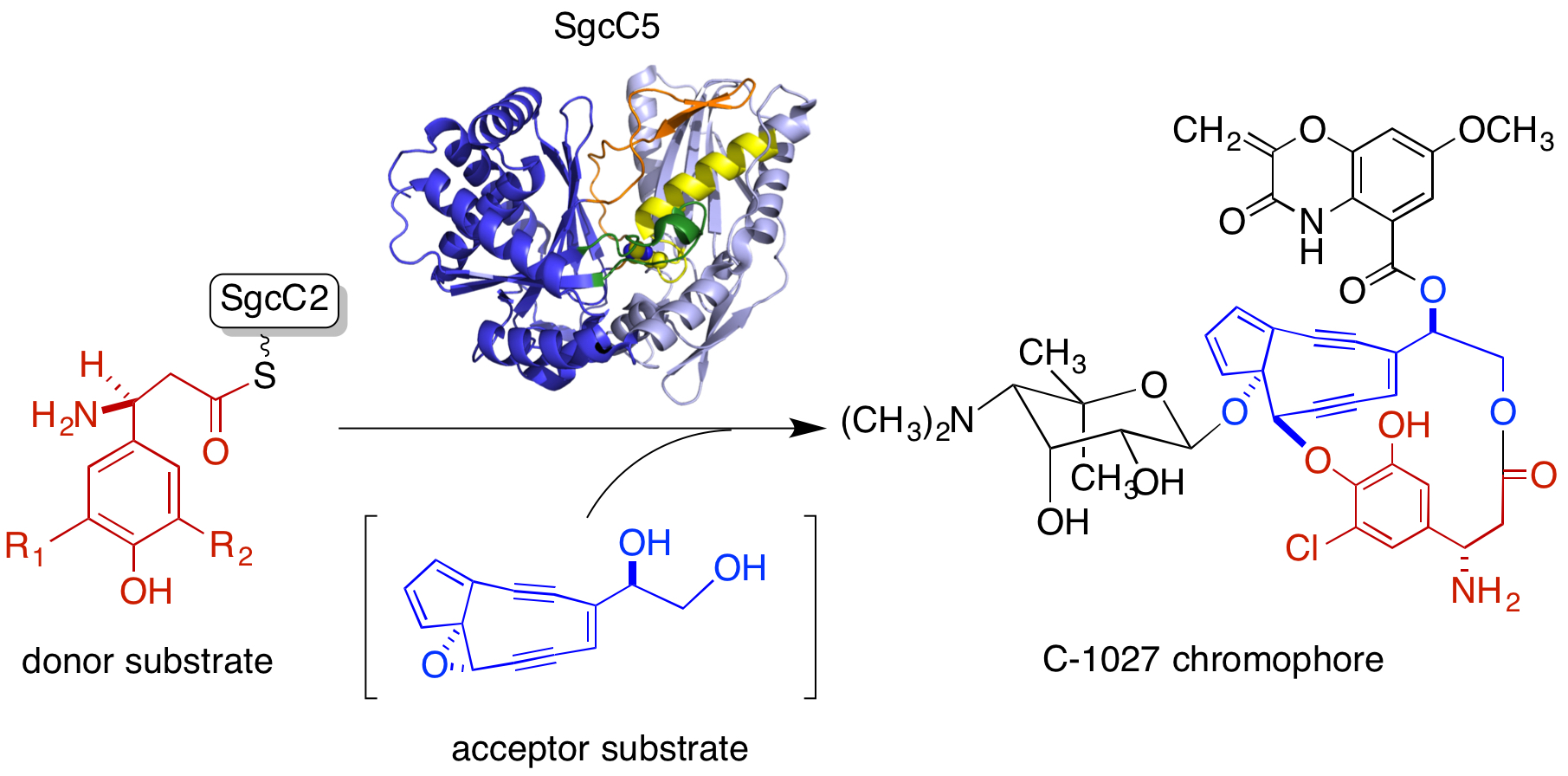
27. Chang, C.-Y.; Lohman, J. R.; Huang T.; Michalska, K.; Bigelow, L.; Rudolf, J. D.; Jedrzejczak, R.; Yan, Y.; Ma, M.; Babnigg, G.; Joachimiak, A.; Phillips Jr., G. N.; Shen, B.* Structural insights into the free-standing condensation enzyme SgcC5 catalyzing ester-bond formation in the biosynthesis of the enediyne antitumor antibiotic C-1027. Biochemistry 2018, 57, 3278–3288.
Read the paper!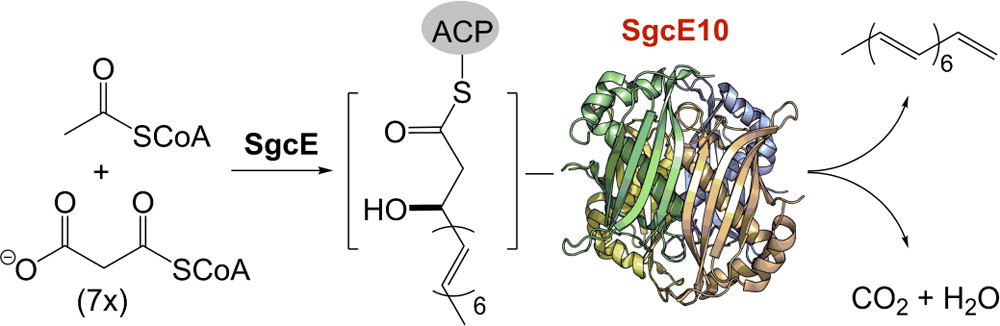
26. Annaval, T.; Rudolf, J. D.; Chang, C.-Y.; Lohman, J. R.; Kim, Y.; Bigelow, L.; Jedrzejczak, R.; Babnigg, G.; Joachimiak, A.; Phillips Jr., G. N.; Shen, B.* Crystal structure of the thioesterase SgcE10 supporting common polyene intermediates in 9- and 10-membered enediyne core biosynthesis. ACS Omega. 2017, 2, 5159–5169.
Read the paper!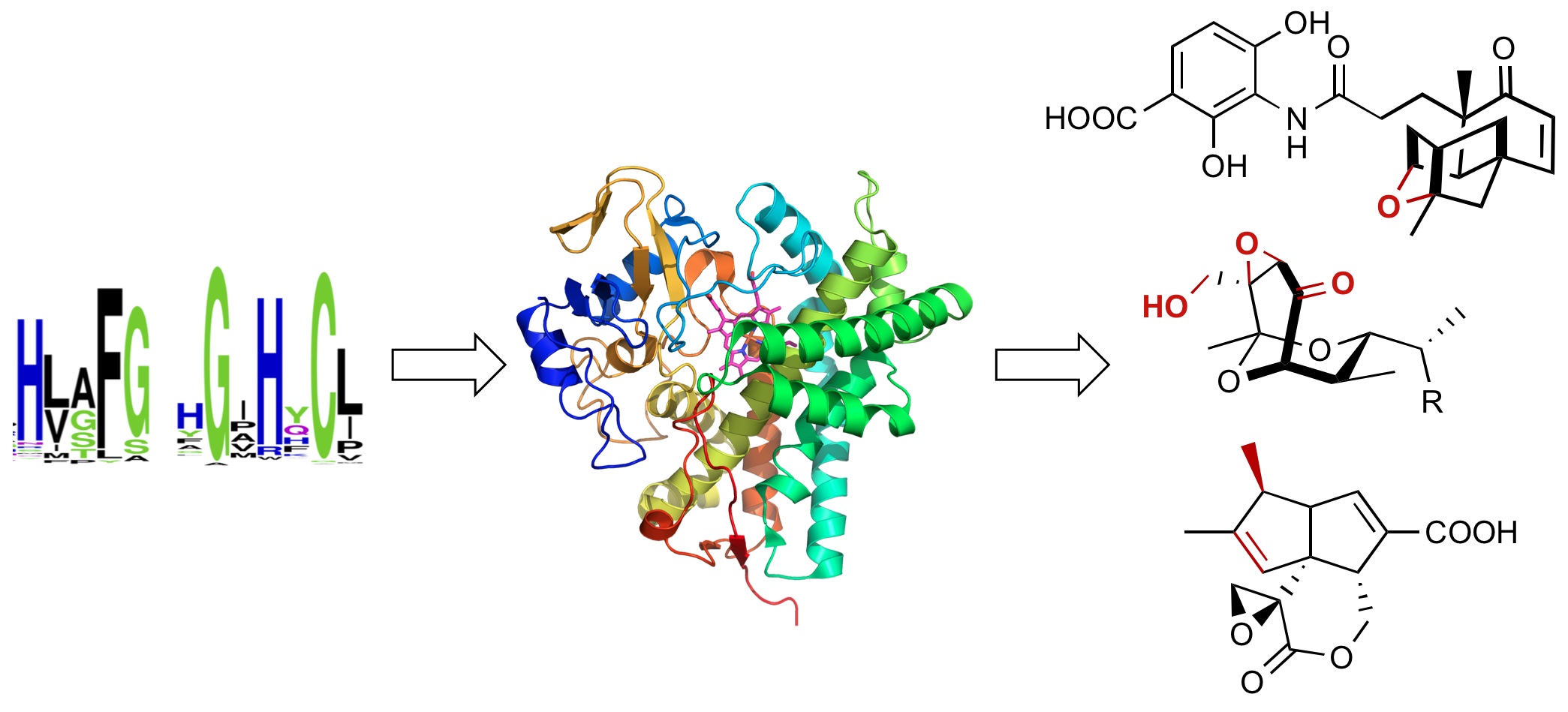
25. Rudolf, J. D.; Chang, C.-Y.; Ma, M.; Shen, B.* Cytochromes P450 for natural product biosynthesis in Streptomyces: sequence, structure, and function. Nat. Prod. Rep. 2017, 34, 1141–1172.
Read the paper!
24. Rudolf, J. D.; Crnovcic, I.; Shen, B.* The role of combinatorial biosynthesis in natural products discovery. In Chemical Biology of Natural Products. Newman, D.J.; Cragg, G.M.; Grothaus, P.G.; Eds. 2017, pp. 87–125.
Read the paper!
23. Falzone, M.; Crespo, E.; Jones, K.; Khan, G.; Korn, V.; Patel, A.; Patel, M.; Patel, K.; Perkins, C.; Siddiqui, S.; Stenger, A.; Yu, E.; Gelber, M.; Scheffler, R.; Nayda, V.; Ravin, A.; Komal, R.; Rudolf, J. D.; Shen, B.; Gullo, V.; Demain, A.* Nutritional control of antibiotic production by Streptomyces platensis MA7327: importance of L-aspartic acid. J. Antibiot. 2017, 70, 828–831.
Read the paper!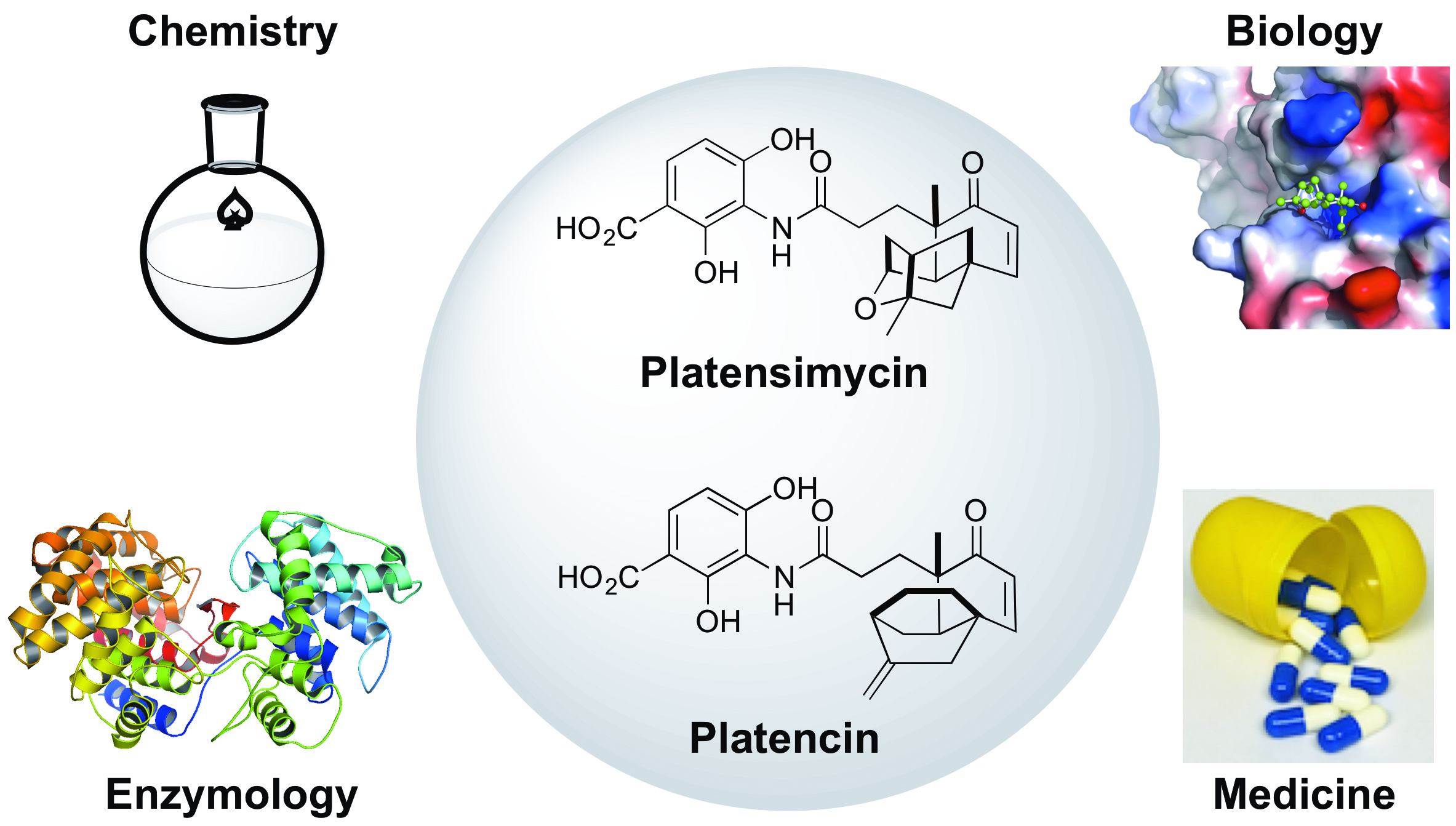
22. Rudolf, J. D.; Dong, L.-B.; Shen, B.* Platensimycin and platencin: Inspirations for chemistry, biology, enzymology, and medicine. Biochem. Pharm. 2017, 133, 139–151.
Read the paper!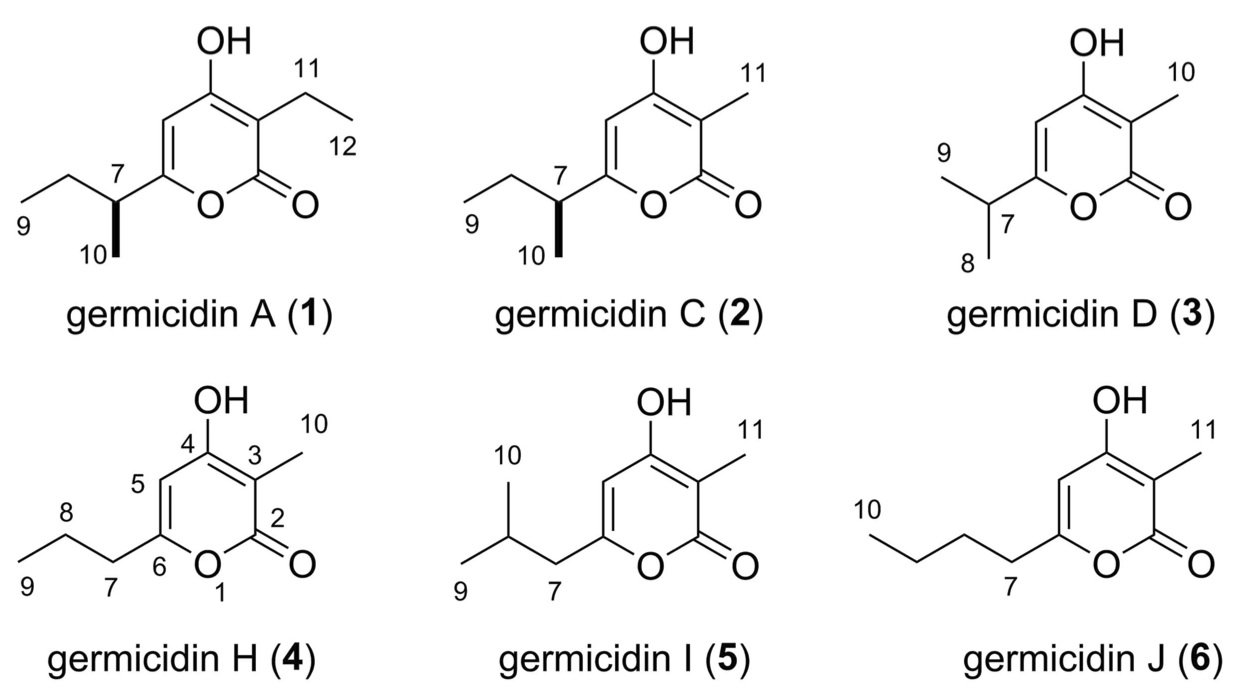
21. Ma, M.; Rateb, M. E.; Yang, D.; Rudolf, J. D.; Zhu, X.; Huang, Y.; Zhao, L.-X.; Jiang, Y.; Duan, Y.; Shen, B.* Germicidins H–J from Streptomyces sp. CB00361. J. Antibiotic. 2017, 70, 200–203.
Read the paper!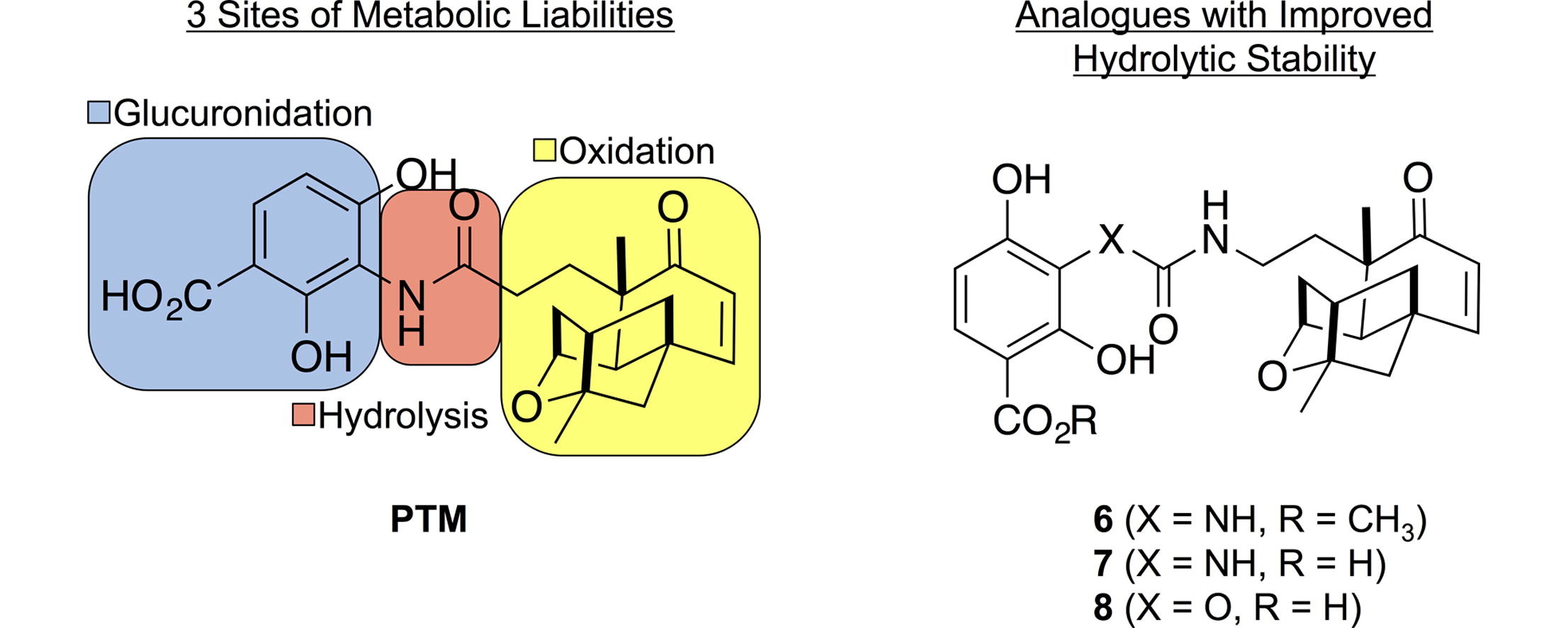
20. Dong, L.-B.; Rudolf, J. D.; Lin, L.; Ruiz, C.; Cameron, M .D.; Shen, B.* In vivo instability of platensimycin and platencin: Synthesis and biological evaluation of urea- and carbamate-platensimycin. Bioorg. Med. Chem. 2017, 25, 1990–1996.
Read the paper!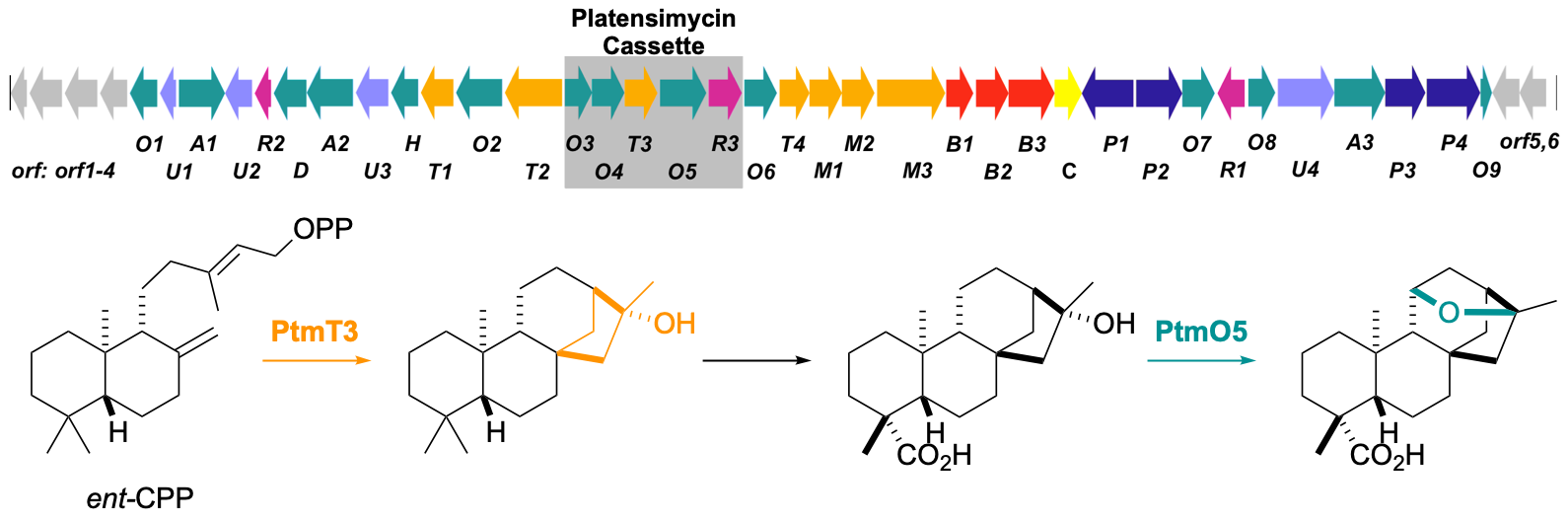
19. Rudolf, J. D.; Dong, L.-B.; Manoogian, K.; Shen, B.* Biosynthetic origin of the ether ring in platensimycin. J. Am. Chem. Soc. 2016, 138, 16711–16721.
Read the paper!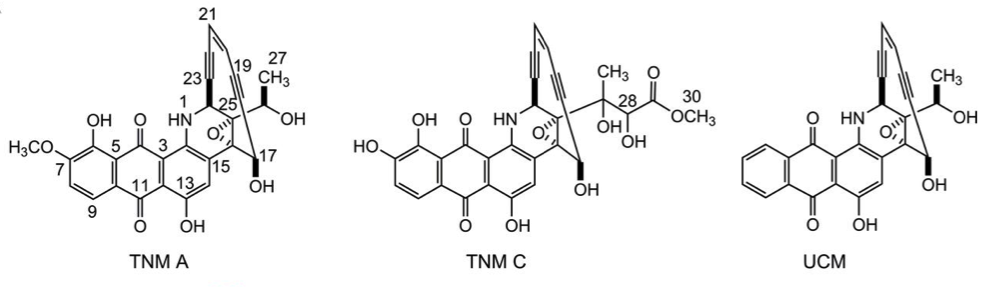
18. Yan, X.; Ge, H.; Huang, T.; Hindra; Yang, D.; Teng, Q.; Crnovcic, I.; Li, X.; Rudolf, J. D.; Lohman, J. R.; Gansemans, Y.; Zhu, X.; Huang, Y.; Zhao, L.-X.; Jiang, Y.; Van Nieuwerburgh, F.; Rader, C.; Duan, Y.; Shen, B.* Strain prioritization and genome mining for enediyne natural products. mBio 2016, 7, e02104-16.
Read the paper!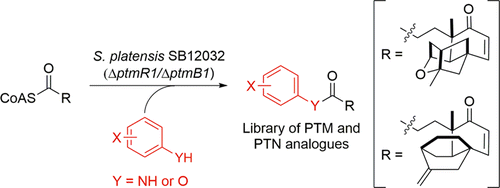
17. Dong, L.-B.; Rudolf, J. D.; Shen, B.* A mutasynthetic library of platensimycin and platencin analogues. Org. Lett. 2016, 18, 4606–4609.
Read the paper!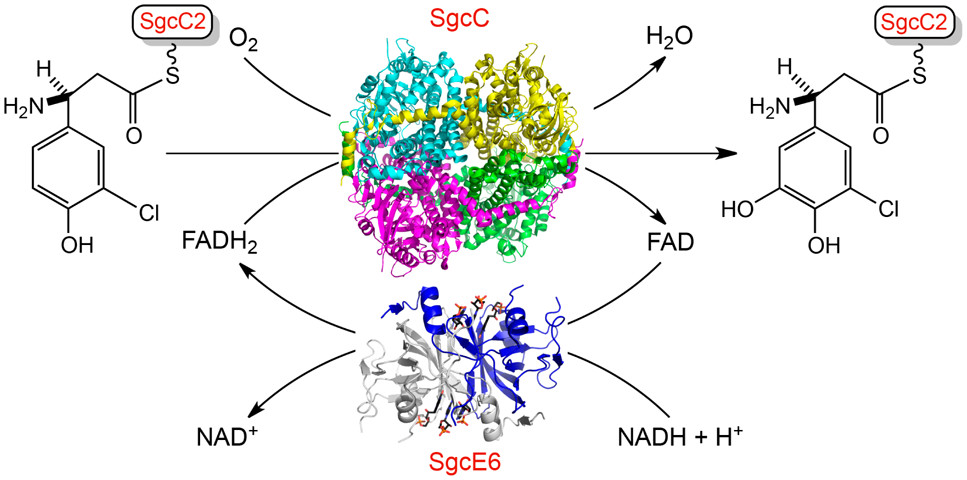
16. Chang, C.-Y.; Lohman, J. R.; Cao, H.; Tan, K.; Rudolf, J. D.; Ma, M.; Xu, W.; Bingman, C. A.; Yennamalli, R. M.; Bigelow, L.; Babnigg, G.; Yan, X.; Joachimiak, A.; Phillips Jr., G.N.; Shen, B.* Crystal structures of SgcE6 and SgcC, the two-component monooxygenase that catalyzes hydroxylation of a carrier protein-tethered substrate during the biosynthesis of the enediyne antitumor antibiotic C-1027 in Streptomyces globisporus. Biochemistry 2016, 55, 5142–5154.
Read the paper!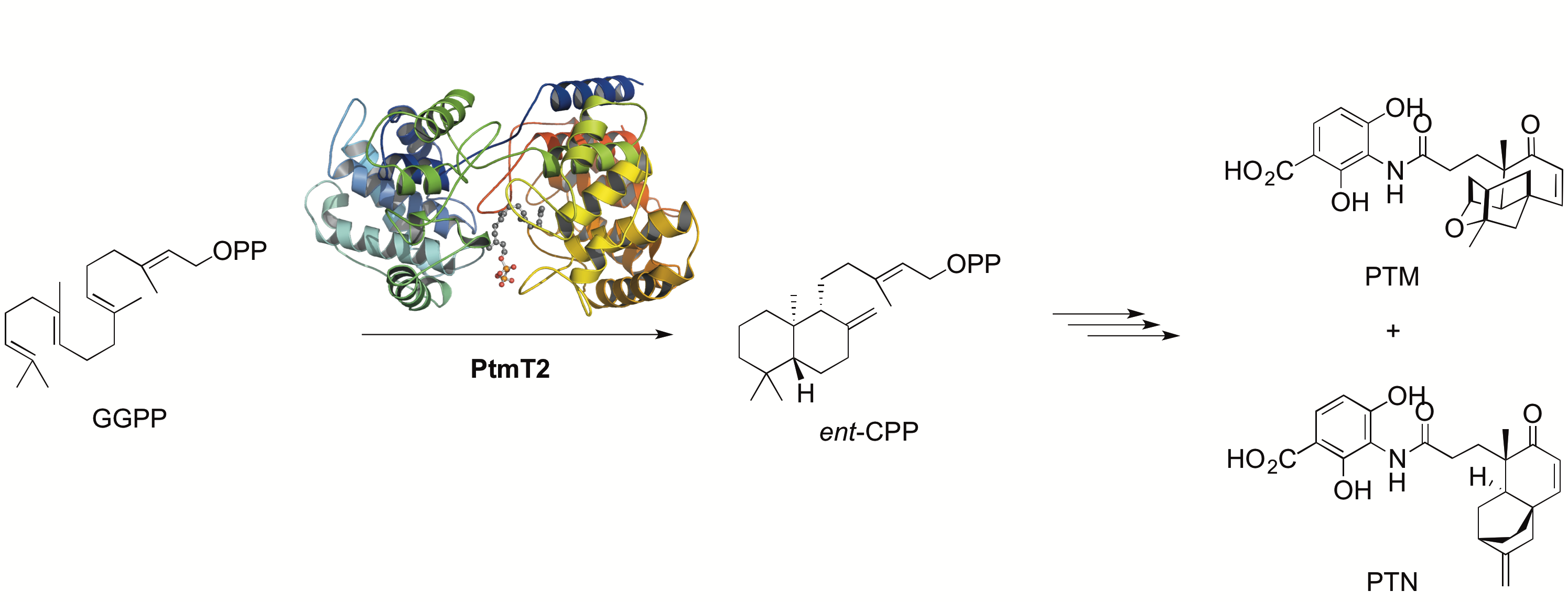
15. Rudolf, J. D.; Dong, L.-B.; Cao, H.; Hatzos-Skintges, C.; Osipiuk, J.; Endres, M.; Chang, C.-Y.; Ma, M.; Babnigg, G.; Joachimiak, A.; Phillips Jr., G.N.; Shen, B.* Structure of the ent-copalyl diphosphate synthase PtmT2 from Streptomyces platensis CB00739, a bacterial type II diterpene synthase. J. Am. Chem. Soc. 2016, 138, 10905–10915.
Read the paper!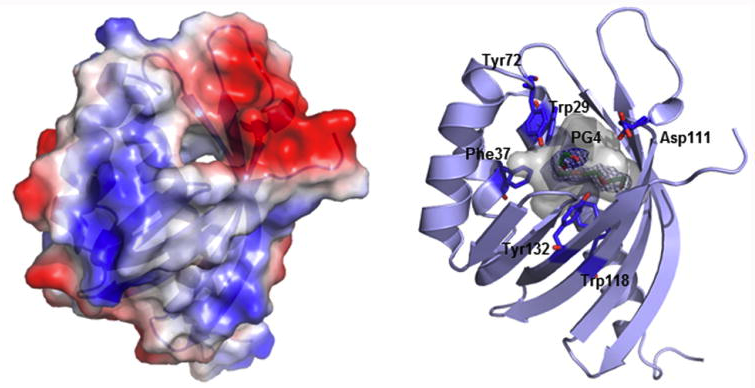
14. Huang, T.; Chang, C.-Y.; Lohman, J. R.; Rudolf, J. D.; Kim, Y.; Chang, C.; Yang, D.; Ma, M.; Yan, X.; Crnovcic, I.; Bigelow, L.; Clancy, S.; Bingman, C. A.; Yennamalli, R. M.; Babnigg, G.; Joachimiak, A.; Phillips Jr., G. N.; Shen, B.* Crystal structure of SgcJ, an NTF2-like superfamily protein involved in biosynthesis of the nine-membered enediyne antitumor antibiotic C-1027. J. Antibiot. 2016, 69, 731–740.
Read the paper!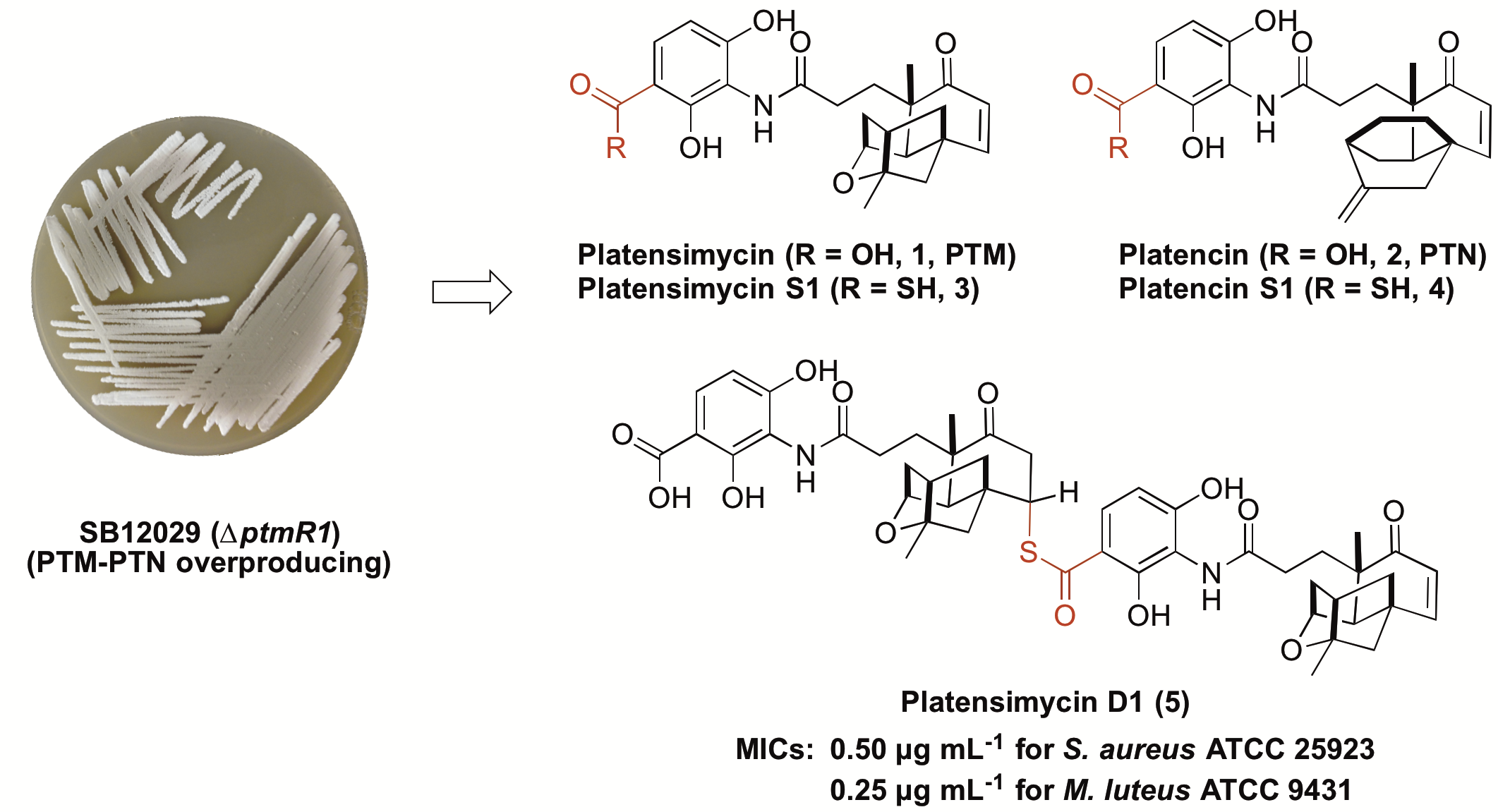
13. Dong, L.-B.; Rudolf, J. D.; Shen, B.* Antibacterial sulfur-containing platensimycin and platencin congeners from Streptomyces platensis SB12029. Bioorg. Med. Chem. 2016, 24, 6348–6353.
Read the paper!
12. Rudolf, J. D.; Yan, X.; Shen, B.* Genome neighborhood network reveals insights into enediyne biosynthesis and facilitates prediction and prioritization for discovery. J. Ind. Microbiol. Biotechnol. 2016, 43, 261–276.
Read the paper!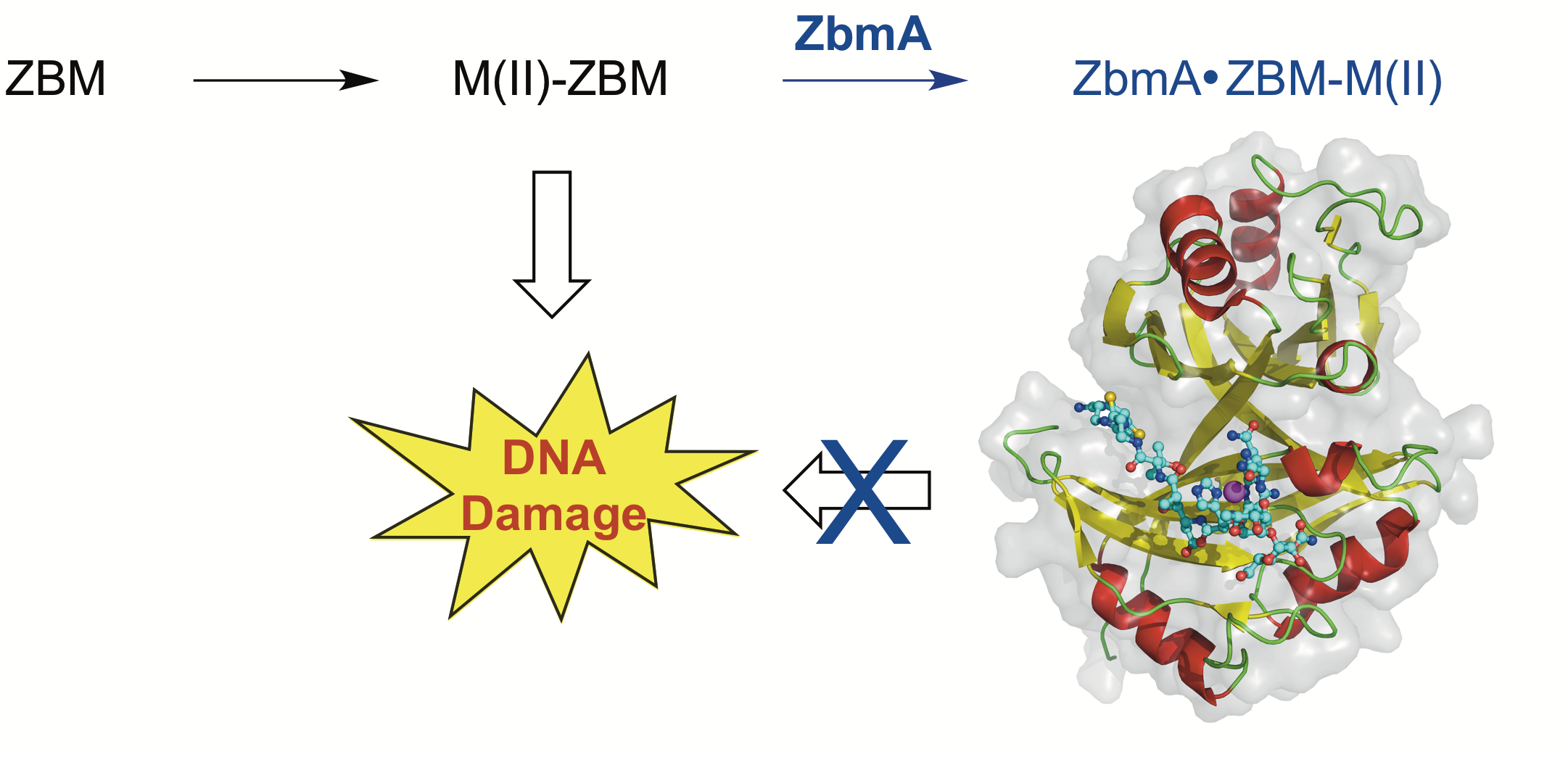
11. Rudolf, J. D.; Bigelow, L.; Chang, C.; Cuff, M. E.; Lohman, J. R.; Chang, C.-Y.; Ma, M.; Yang, D.; Clancy, S.; Babnigg, G.; Joachimiak, A.; Phillips Jr., G. N.; Shen, B.* Crystal structure of the zorbamycin-binding protein ZbmA, the primary self-resistance element in Streptomycesflavoviridis ATCC21892. Biochemistry 2015, 54, 6842–6851.
Read the paper!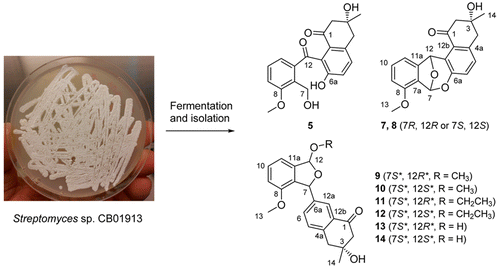
10. Ma, M.; Rateb, M.; Teng, Q.; Yang, D.; Rudolf, J. D.; Zhu, X.; Huang, Y.; Zhao, L.-X.; Jiang, Y.; Li, X.; Rader, C.; Duan Y.; Shen, B.* Angucyclines and angucyclinones from Streptomyces sp. CB01913 featuring C-ring cleavage and expansion. J. Nat. Prod. 2015, 78, 2471–2780.
Read the paper!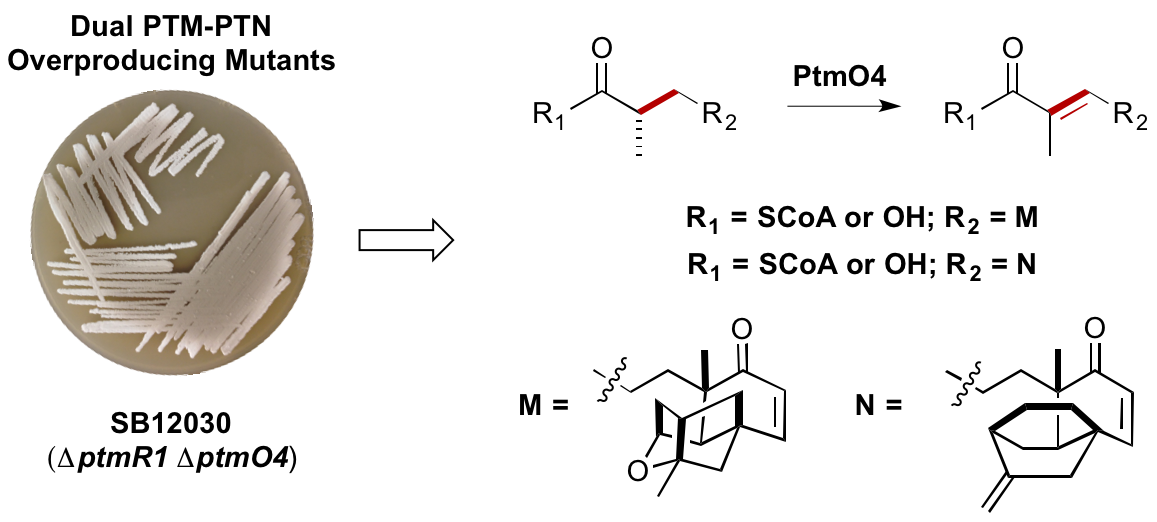
9. Rudolf, J. D.; Dong, L.-B.; Huang, T.; Shen, B.* A genetically amenable platensimycin- and platencin-overproducer as a platform for biosynthetic explorations: a showcase of PtmO4, a long-chain acyl-CoA dehydrogenase. Mol. BioSyst. 2015, 11, 2717–2726.
Read the paper!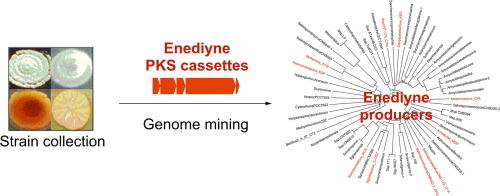
8. Shen, B.;* Hindra; Yan, X.; Huang, T.; Ge, H.; Yang, D.; Qihui, T.; Rudolf, J. D.; Lohman, J. R. Enediynes: Exploration of microbial genomics to discover new anticancer drug leads. Bioorg. Med. Chem. 2015, 25, 9–15.
Read the paper!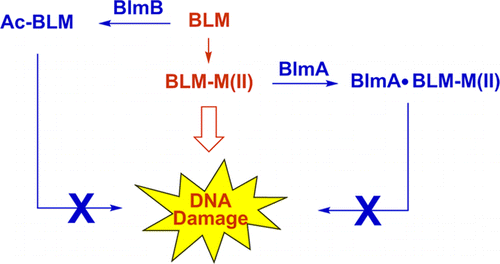
7. Coughlin, J. M.; Rudolf, J. D.; Wang, L.; Galm, U.; Wendt-Pienkowski, E.; Tao, M.; Shen, B.* BlmB and TlmB provide resistance to the bleomycin family of antitumor antibiotics by N-acetylating metal-free bleomycin, tallysomycin, phleomycin, and zorbamycin. Biochemistry 2014, 53, 6901–6909.
Read the paper!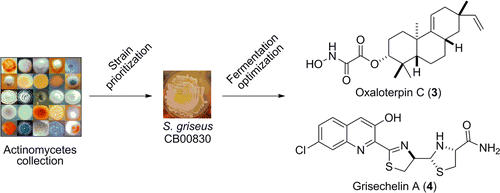
6. Hindra; Huang, T.; Yang, D.; Rudolf, J. D.; Xie, P.; Xie, G.; Teng, Q.; Lohman, J. R.; Zhu, X.; Huang, Y.; Zhao, L.-X.; Jiang, Y.; Duan, Y.; Shen, B.* Strain prioritization for natural product discovery by a high-throughput real-time PCR method. J. Nat. Prod. 2014, 77, 2296–2303.
Read the paper!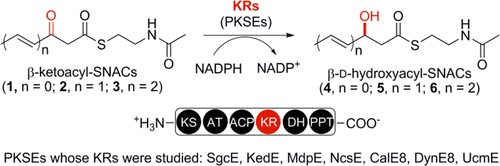
5. Ge, H.-M.; Huang, T.; Rudolf, J. D.; Lohman, J. R.; Huang, S.-X.; Guo, X.; Shen, B.* Enediyne polyketide synthases stereospecifically reduce the β-ketoacyl intermediates to β-D-hydroxyacyl intermediates in enediyne core biosynthesis. Org. Lett. 2014, 16, 3958–3961.
Read the paper!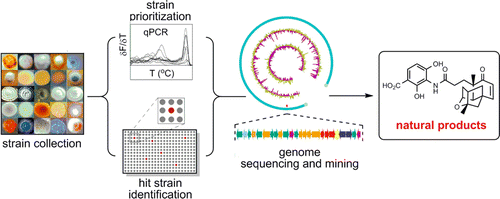
4. Xie, P.; Ma, M.; Rateb, M.; Shaaban, K.; Yu, Z.; Huang, S.-X.; Zhao, L.-X.; Zhu, X.; Yan, Y.; Peterson, R. M.; Lohman, J. R.; Yang, D.; Yin, M.; Rudolf, J. D.; Jiang, Y.; Duan, Y.; Shen, B.* Biosynthetic potential-based strain prioritization for natural product discovery – a showcase for diterpenoid producing actinomycetes. J. Nat. Prod. 2014, 77, 377–387.
Read the paper!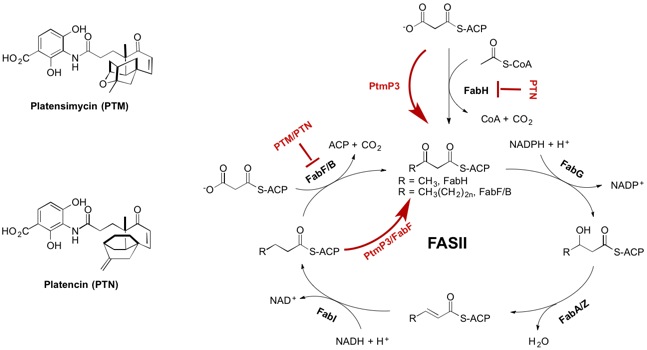
3. Peterson, R. M.; Huang, T.; Rudolf, J. D.; Smanski, M. J.; Shen B.* Mechanisms of self-resistance in the platensimycin and platencin producing Streptomyces platensis MA7327 and MA7339 strains. Chem. Biol. 2014, 21, 389–397.
Read the paper!University of Utah – Graduate School
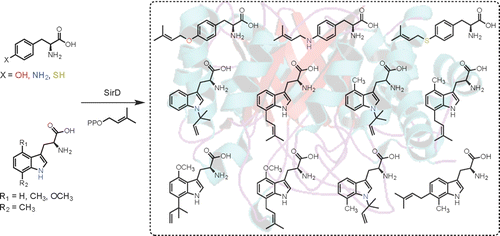
2. Rudolf, J. D.; Poulter, C. D.* Tyrosine O-prenyltransferase SirD catalyzes S-, C-, and N-prenylations on tyrosine and tryptophan derivatives. ACS Chem. Biol. 2013, 8, 2707–2714.
Read the paper!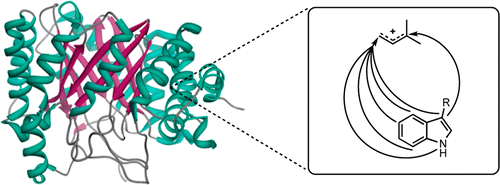
1. Rudolf, J. D.; Wang, H.; Poulter, C. D.* Multisite prenylation of 4-substituted tryptophans by dimethylallyltryptophan synthase. J. Am. Chem. Soc. 2013, 135, 1895–1902.
Read the paper!70
Publications
2510
Citations
28
h-index
